Genomics of Early Growth and Cardio-Metabolic Health

- Fasil Tekola-Ayele,
PhD, Senior Investigator, Epidemiology Branch, DiPHR - Kunal Kathuria, PhD, Staff Scientist
- Tesfa Habtewold, PhD, Postdoctoral Visiting Fellow
- Prabhavi Wijesiriwardhana, PhD, Postdoctoral Visiting Fellow
- Shila Deljookorani, BS, Postbaccalaureate Fellow Undergraduate Scholars Program
The early developmental period marks an important milestone for lifelong health, including later-onset cardiovascular, metabolic, and neuropsychiatric diseases. Understanding the complex genetic and environmental factors that underlie these relationships is crucial for guiding the development of early preventive and therapeutic interventions for maximizing health across the life span. In this context, the placenta, which is a functionally critical transient organ during pregnancy between the maternal and fetal interface, is increasingly implicated not just in pregnancy maintenance and fetal development regulation but also in early origins of adult-onset diseases. Investigation of genetic and prenatal environmental factors that alter the epigenetic profile of the placenta can unlock biological underpinnings of physiologic traits and disease risk. Such advances will provide early intervention opportunities for cardio-metabolic outcomes. Absence of the placenta in many genomics databases has hindered functional advance. Moreover, perinatal genomic studies have not adequately covered the genetic variation landscape of human populations, which impedes biological understanding of diseases and could undermine population-wide benefits of genomic-informed health care. In the genetic epidemiology research group, we study genetic mechanisms in early growth variations at the maternal-placental-fetal interface and their links with cardio-metabolic outcomes in multi-ancestral human populations.
Maternal genetic influence on fetal growth and interactions with non-genetic factors
Genome-wide association studies (GWAS) have identified maternal and fetal genetic variants associated with birth weight in European ancestry populations. However, the genetic architecture of longitudinal fetal growth may not overlap with that of birth weight. Birth weight does not represent the trajectory of fetal growth throughout pregnancy. Two fetuses with different sizes at a given gestational age may arrive at the same size at birth although undergoing different growth rates. Fetal growth rate has clinical values, such as improved prediction of small for gestational age, placental insufficiency, and fetal death, that cannot be obtained from a one-time fetal size measure. We previously found that the contribution of genetics to fetal growth relative to environmental factors varies throughout pregnancy, with a peak observed at end of the second trimester and the early third trimester. The trajectory of fetal growth varies with the genome-wide average levels of maternal genetic ancestry. Genetic regions with higher levels of African and Amerindigenous ancestry have been associated with fetal growth measures, pointing to molecular pathways involved in fetal growth regulation. We are investigating genome-wide maternal loci to identify genetic variants associated with the longitudinal trajectory of fetal weight throughout pregnancy by combining datasets of pregnancy cohorts. Using a meta-regression approach that accommodates genetic ancestry-related allelic-effect heterogeneity, we identified novel genetic variants associated with change in fetal weight trajectory. For most of the identified loci, allelic-effect heterogeneity was significantly correlated with genetic ancestry.
The maternal genome interacts with various non-genetic factors, including obstetric conditions, in influencing fetal growth. We found that maternal genetic ancestry distance (the similarity of an individual’s genome to a given population genome reference) modifies the influence of maternal pre-pregnancy obesity on placental and neonatal weights. Moreover, multiparity (having a prior pregnancy lasting more than 20 weeks of gestation) may buffer the birthweight-reducing effect of the maternal genome.
Prenatal glycemic and oxidative cumulative exposures linked to placental aging and omics
The placenta offers an opportunity to study the molecular pathways targeted by prenatal environmental exposures, interactions between genetic and environmental factors, and maternal-fetal adaptive mechanisms. Maternal cardio-metabolic status during pregnancy influences the success of pregnancy as well as long-term child and maternal health outcomes. Placental molecular features can be altered by maternal cardio-metabolic dysregulations. Therefore, epigenetic mechanisms in the placenta may be a potential link between maternal cardio-metabolic status, fetal growth, and offsprings' future risk of cardio-metabolic diseases. Accelerated aging of the placenta can reduce its functional capacity and is implicated in adverse pregnancy outcomes, including fetal growth restriction, preeclampsia, and preterm birth. Using the concept of “epigenetic clocks,” placental epigenetic age acceleration can be determined as the difference between the epigenetic and gestational age of the placenta, and indicates the biological age of the placenta relative to its chronological gestational age. We previously found that accelerated placental epigenetic aging is associated with fetal growth in a sex-dependent manner. Maternal obesity, gestational weight gain, and dyslipidemia are associated with placental epigenetic age acceleration. These studies highlight the fact that the placental “clock” can be an avenue to study the molecular etiologies of fetal growth abnormalities, and can act as a biomarker for tracking prenatal environmental and genetic predisposition.
Placental response to prenatal exposures can vary based on gestational timepoints during pregnancy. Therefore, epigenetic, gene expression, and aging “clock” profiles of placentas may track prenatal exposures at specific gestational weeks as well as cumulative exposures over gestation. This line of investigation is often constrained by limited datasets with exposures measured longitudinally across pregnancy. We measured glycemic, oxidative stress, and inflammation markers in maternal plasma at gestational weeks 8–13, 16–22, 24–29, and 34–37 in the NICHD Fetal Growth Studies. The lab investigated the association between multiple measures of glycemic markers (glucose, HbA1c, insulin) and placental age acceleration. Cumulative levels of each glycemic marker by trimester were estimated using the area under the curve. Elevated glucose, HbA1c, and insulin cumulative levels during second trimester were associated with lower placental age acceleration. This finding is aligned with histopathologic placental villous immaturity observed in hyperglycemic pregnancies. Developmental immaturity of the placenta is associated with fetal death and delayed child neurodevelopment, conditions more common in hyperglycemic pregnancies. Therefore, hyperglycemia may alter the placental “clock” leading to placental developmental immaturity, which, in turn, has long-term health effects. We are investigating the link between longitudinally measured maternal plasma oxidative stress and inflammation markers and placental age acceleration.
Fetal sex, placental omics, and neonatal/adult health outcomes
Sex differences have been documented in fetal growth and several childhood and adult traits and diseases. Sex differences in phenotypes may in part have prenatal molecular origins embedded in the placenta, including sex-dependent transcriptomic.htmlepigenetic profiles and response to prenatal exposures. Genetic modulation of placental function can also vary by fetal sex. Therefore, understanding sex-dependent genetic regulation of methylation and gene expression in the placenta can inform mechanisms that underlie pregnancy outcomes and early origins of sex differences in numerous phenotypes. Our previous work showed that sex differences exist in the relations between placental epigenetic aging and fetal growth. Placental gene expression linked with increased odds of small-for-gestational age neonates involves pathways that differ between male and female fetuses. Maternal cardio-metabolic and psychosocial influences on the placental epigenome also vary by fetal sex.
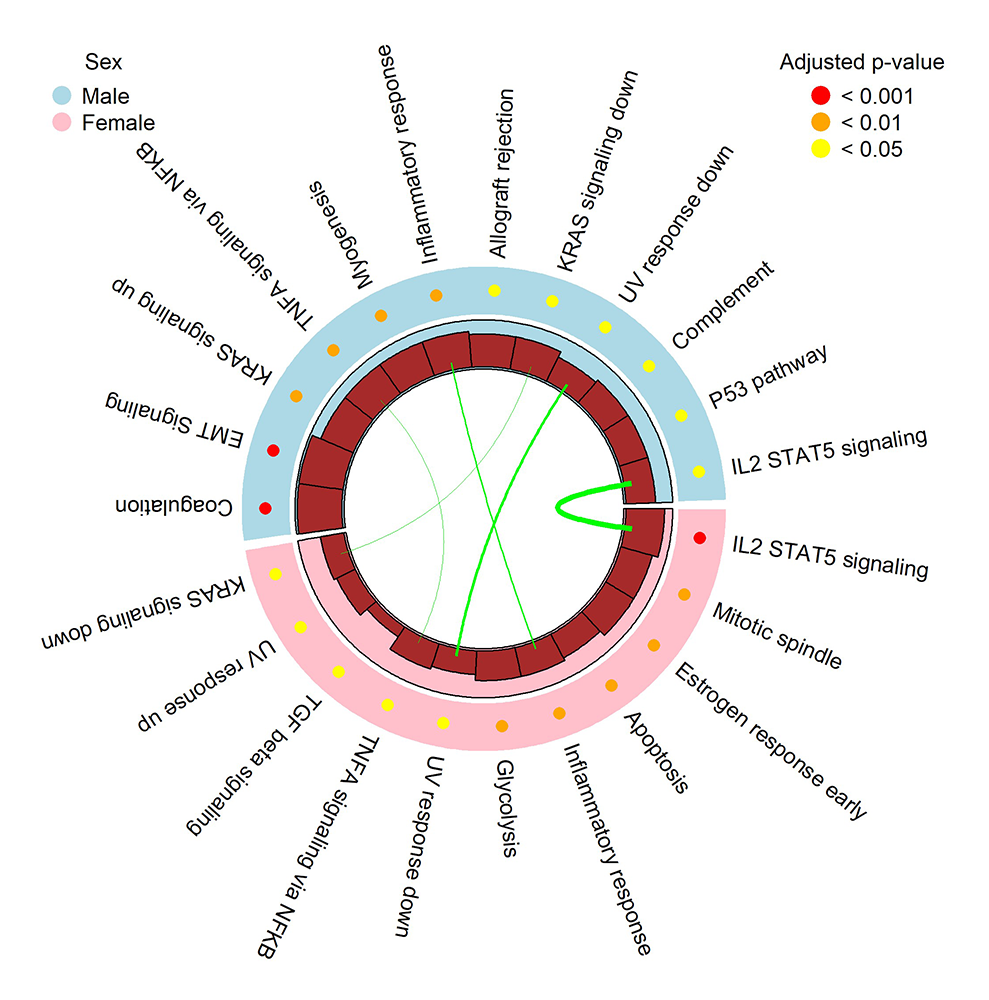
Our lab conducted a study to uncover how intersections between fetal sex, placental gene expression, and methylation levels and their genetic regulation underpin fetal growth outcomes and adult diseases. Key highlights of the study include the following. 1) Extensive sex differences exist for placental methylation and sex-dependent genetic regulation of methylation. However, limited sex differences were found for both the level and the genetic regulation of placental autosomal gene expression. Downstream analyses found largely distinct sex-dependent placental methylation, gene expression, and their genetic regulation. 2) Several sex-differentiated genomic loci converged at important molecular processes involved in health and disease. While male-hypermethylation was more likely to be associated with increased neonatal size, female-hypermethylation was linked to increased placental weight. Sex-differentiated genes were enriched for several hallmark gene sets, which represent multiple coherently expressed genes in a well defined biological process. Male-biased placental methylation was enriched for the hemostasis/coagulation hallmark gene set (Figure 1), suggesting epigenetically suppressed coagulation as one potential explanation for male-preponderance in some pregnancy complications. Sex-specific genetic effects on methylation and gene expression colocalized with several GWAS loci for birthweight, breast cancer, and allergic diseases, yielding a molecular framework for future investigation of early origins of these health outcomes. 3) Sex-differentiated loci were significantly enriched for human imprinted genes, a parent-of-origin-dependent gene expression regulation critical for placental function and fetal development. This observation suggests that “fetal sex-imprinting crosstalk” may act as a molecular balance to prevent uncontrolled aberrant growth.
Figure 1. Hallmark gene sets significantly enriched in males and females
Outer track: enriched hallmark gene sets; middle track: adjusted p-value; orange bars in innermost track: number of overlapping genes between males and females; green lines: hallmark gene sets shared between males and females, with width proportional to number of shared genes. Source: Reference 1.
Figure 1. Hallmark gene sets significantly enriched in males and females
Outer track: enriched hallmark gene sets; middle track: adjusted p-value; orange bars in innermost track: number of overlapping genes between males and females; green lines: hallmark gene sets shared between males and females, with width proportional to number of shared genes. Source: Reference 1.
Overall, the study highlighted that the placenta is actively involved in setting the developmental foundation of pregnancy complications and future health outcomes shaped by fetal sex. The study yielded a resource catalog of sex-differentiated methylation, gene expression, and their genetic regulation in human placenta. In an ongoing study, we are investigating sex differences in placental gene expression within placental cell type clusters, how these differences may evolve as a function of gestational age, and the implications on pregnancy outcomes.
Insights into fetal growth regulation using human placental multi-omics
Placental epigenome, transcriptome, and its epigenetic age acceleration are under substantial genetic control. The implicated genes and genetic mechanisms that link placental age acceleration and fetal growth are yet to be identified. Elucidating the pathophysiologic links between placental epigenetic aging and fetal growth outcomes can increase the clinical utility of the placental epigenetic aging clock in predicting adverse pregnancy outcomes. Placental epigenetic acceleration has a high heritability, suggesting substantial genetic control; however, no prior studies have investigated its genetic basis. To address these research gaps, we initiated a new study as part of the Study of Pregnancy and Neonatal Health (SPAN) to establish a prospective pregnancy cohort of multi-ancestral pregnant women in the gGAP (genomics in growth and placenta) aim. gGAP will integrate multi-omics approaches (genome-wide genotypes, placental methylation, placental gene expression) to identify and functionally characterize genetic mechanisms that underpin placental epigenetic aging, longitudinal fetal growth, and the links between the two in a multi-ancestral cohort oversampled by African Americans. The goals of gGAP in SPAN are to: (1) investigate the fetal genome’s contribution in the genetic architecture of fetal growth for which data are lacking; (2) interrogate the effect on fetal growth of gene-environment interactions in the context of maternal psychosocial, obstetric, and cardio-metabolic factors; (3) refine causal loci by leveraging genomic variation of multi-ancestral populations and build a perinatal multi-omics resource; and (4) integrate genomics with placental imaging, histopathology, and proteomics for an in-depth mechanistic investigation of fetal growth and other pregnancy outcomes. At baseline (less than 20 gestational weeks), pregnant women complete a detailed assessment using questionnaire items and physical examination, and provide blood, saliva, and urine samples. During third trimester, fetal biometry ultrasound examination and blood collection are undertaken. At delivery, cord blood and placental biopsy tissue are collected, gross examination and photographic image of the placenta are taken, and neonatal anthropometry, body composition, and neuro-behavioral measures are obtained. Detailed chart abstraction is completed including first- and second-trimester fetal biometry obtained by clinical ultrasound. SPAN data collection is ongoing at four clinical sites across the United States. Two new clinical sites have been added this year.
Genomics of early childhood longitudinal growth/obesity
A key motivation for our lab's research is to understand the mechanistic basis of the relationship between early-life growth and later-life cardio-metabolic health. The lab’s prior work, as well as that of others, hinted at genetic effects shared between birthweight and child and adult growth measures and cardiometabolic outcomes. Genetic factors with overlapping effects on birthweight and cardio-metabolic diseases in later life may partly explain early origins of later-life health, complementing other mechanisms such as in utero programming. Such advances are important for developing molecular interventions to address pressing public health problems, including childhood obesity. Childhood body mass index (BMI) differences result from genetic and environmental influences. Heritability of childhood BMI has been estimated to be 40–70%. Therefore, identifying genetic factors that regulate BMI in early childhood, including genes with age-specific effects, can offer a window of opportunity for early diagnosis and for development of therapeutics for obesity and related diseases. However, genetic factors that influence early childhood BMI, in particular during infancy and prior to adiposity rebound, are not well understood. Longitudinal cohorts with repeatedly measured anthropometry measures are crucial to understanding shared as well as age-specific genetic mechanisms involved in growth and metabolic phenotypes.
To address these research gaps, cohorts with well characterized longitudinal phenotypes and multi-ancestry genomic data are needed. We initiated a new genetic study embedded in the collaborative perinatal project (CPP). The CPP is a prospective pregnancy cohort that enrolled more than 48,000 pregnant women (46% White, 46% African American, 7% Puerto Rican, 1% others) at 12 medical centers in the United States in 1959–1965. Women were followed during pregnancy. Children were monitored at seven consecutive times that cover important developmental milestones from birth through early school ages (birth, 4, 8, and 12 months and 3, 4, and 7 years). Through a pilot study, we optimized approaches to achieve high quality and reproducible genotyping of DNA from CPP–stored biospecimens. Building on the success of the pilot, we set out to establish a genomic database for children in the CPP, with the following goals: (1) to establish a genomic database for children in the CPP cohort (n=10,800; 5,400 African American and 5,400 European American); (2) to identify genetic influences on early growth anthropometry and obesity-related phenotypes at seven longitudinal time points from birth through school age, using multi-ancestral and admixture mapping approaches; and (3) to identify molecular pathways in the link between fetal and early childhood growth and developmental phenotypes. The study will facilitate insights into early origins of adult diseases and interventions by determining genetic effects that are transient and those that are sustained across ages. The CPP cohort enrolled participants prior to lifestyle changes that led to the obesity epidemic. This lack of major obesogenic environment is advantageous to identify genes whose effect on obesity is largely genetic. The potential interaction between genetic factors discovered via the CPP and recent environmental modifiers will be evaluated in other recent cohorts. This year marks successful completion of DNA extraction and genotyping for more than 9,000 samples. Genetic data quality processing is ongoing.
Additional Funding
- Undergraduate Scholarship Program awarded to Shila Deljookorani
- NICHD Director’s Strategic Planning grant to Fasil Tekola-Ayele
Publications
- Sex-differentiated placental methylation and gene expression regulation has implications for neonatal traits and adult diseases. Nat Commun 2025 16(1):4004
- Longitudinal maternal glycemia during pregnancy and placental epigenetic age acceleration. Clin Epigenetics 2025 17(1):19
- Maternal obesity and ancestry distance in influencing birth outcomes. Int J Obes (Lond) 2025 49(8):1532-1542
- Association of placental weight and placental-fetal weight ratio with DNA methylation in placenta. Epigenomics 2025 17(9):589-598
- Maternal parity modifies the association of birthweight polygenic score with fetal growth. Sci Rep 2025 15(1):27915
Collaborators
- Karin Fox, MD, MEd, University of Texas Medical Branch, Galveston, TX
- Brenna Hughes, MD, Duke University, Durham, NC
- Carmen Marsit, PhD, Emory University, Atlanta, GA
- George L. Maxwell, MD, Inova Fairfax Hospital, Falls Church, VA
- Daniel Molina, PharmD, Technical Resources International, Inc., Bethesda, MD
- Robert Silver, MD, University of Utah, Salt Lake City, UT
- Alan Tita, MD, PhD, University of Alabama at Birmingham, Birmingham, AL
- Michael Tsai, PhD, University of Minnesota, Minneapolis-St. Paul, MN
- Ronald Wapner, MD, Columbia University, New York, NY
- Timothy Wen, MD, MPH, University of California San Diego Health, San Diego, CA
Contact
For more information, email ayeleft@mail.nih.gov or visit https://irp.nih.gov/pi/fasil-tekola-ayele.