Quantitative Imaging and Tissue Sciences

- Peter J. Basser,
PhD, Head, Section on Quantitative Imaging and Tissue Sciences - Ferenc Horkay, PhD, Staff Scientist (deceased)
- Teddy Cai, PhD, Postdoctoral Intramural Research Training Award Fellow
- Ella Wilczynski, PhD, Postdoctoral Visiting Fellow
- Julian Rey, PhD, Postdoctoral Research Associate Training (PRAT) Fellow
- Alexandru V. Avram, PhD, Collaborating Contract Scientist
- Rea Ravin, PhD, Collaborating Contract Scientist
- Kadharbatcha Saleem, PhD, Collaborating Contract Scientist
- Mihilka Gangolli, PhD, Collaborating Scientist funded via the Henry Jackson Foundation and the Military Traumatic Brain Injury Initiative
- Michal Komlosh, PhD, Collaborating Scientist funded via the Henry Jackson Foundation and the Military Traumatic Brain Injury Initiative
- Magdoom Kulam, PhD, Collaborating Scientist funded via the Henry Jackson Foundation and the Military Traumatic Brain Injury Initiative
- Nathan Williamson, PhD, Collaborating Scientist funded via the Henry Jackson Foundation and the Military Traumatic Brain Injury Initiative
- R. Douglas Fields, PhD, Special Volunteer
- Iren Horkayne-Szakaly, MD, Special Volunteer
- Shinjini Kundu, MD, PhD, Special Volunteer
In our tissue-sciences research, we are interested in how tissue microstructure, hierarchical organization, composition, and material and transport properties all work together to affect biological function or dysfunction. We investigate biological and physical model systems at various length- and time-scales, performing biophysical measurements and developing novel physical/mathematical models (e.g., continuum models) to explain their functional properties and behavior. We use water to probe tissue structure and function from nanometers to centimeters and from microseconds to lifetimes. Our armamentarium includes small-angle neutron scattering (SANS), small-angle X-ray scattering (SAXS), static light scattering (SLS), dynamic light scattering (DLS), atomic force microscopy (AFM), osmometry, and multi-dimensional nuclear magnetic resonance (NMR) methods. We develop research tools that can be translated from bench-based quantitative methodologies to bedside biomarkers to aid in diagnosis, prognosis assessment, and therapy.
Our work is directed toward making critical ‘invisible’ biological structures and transport processes ‘visible.’ Our quantitative imaging group uses knowledge of physics, engineering, applied mathematics, imaging, and computer sciences, as well as key insights gleaned from our tissue-sciences research, to discover, vet, and develop novel quantitative imaging biomarkers that can detect changes in tissue composition, microstructure, and/or microdynamics with high sensitivity and specificity so as to assess normal and abnormal developmental trajectories, diagnose childhood diseases and disorders, and characterize degeneration and trauma, such as mild traumatic brain injury (mTBI). MRI is our imaging modality of choice because it is non-invasive, non-ionizing, usually requires no exogenous contrast agents or dyes, and is generally deemed safe and effective for use with mothers, their fetuses, and children in both clinical and research settings. Critical to this enterprise is our ability to follow water as it migrates through complex media as a probe of its microstructure, and to assess water's interactions with biomolecules to identify distinct water compartments in tissues.
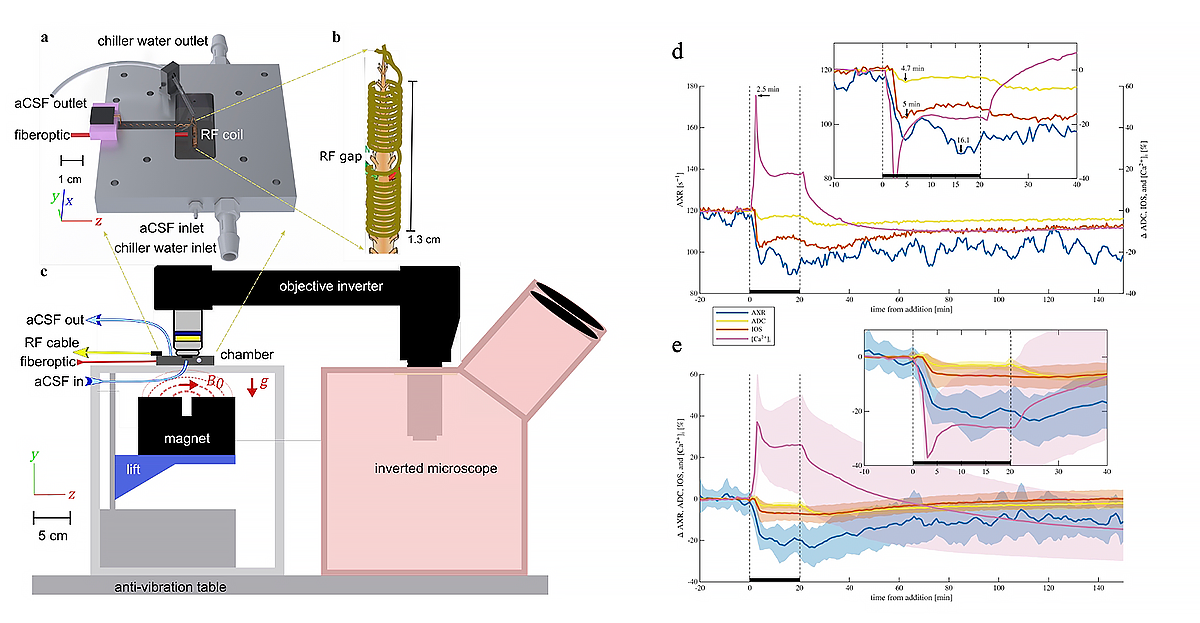
Figure 1. Combined MR profiling and optical imaging system for studying water transport processes in tissue specimen
Test system developed for interrogating and validating quantitative diffusion and exchange MR imaging biomarkers via simultaneous real-time NMR and optical microscopy. a) Setup includes the single-sided magnet (NMR MOUSE, Magritek) and the ex vivo tissue chamber (David Ide, Section on Instrumentation, NIMH), modified 3-D technical drawing of the test chamber. b) Drawing of the solenoid radiofrequency (RF) coil containing a viable mouse spinal cord specimen, showing a gap in the RF coil where the sample is imaged. c) Technical drawing of the experimental setup showing the single-sided permanent magnet projecting a magnetic field on the sample from below and the inverted microscope imaging the sample from above. d) Representative sample. e) mean ± SD of AXR (6-pt moving average), Apparent Diffusion Coefficient (ADC), Intrinsic Optical Signal (IOS), and [Ca2+i] fluorophore signal during 20 min 50 mM KCl washin and washout (n = 12, three of which included [Ca2+i] imaging). Insets show zoomed regions with [Ca2+i] inverted for comparison. Arrows in a) mark signal minima and maxima. The data reveal connections between the metrics and their sensitivity to pathological cellular swelling during depolarization. The study helps facilitate the translation from cellular neurophysiology findings to the clinic, and to provide a framework to validate associated MRI methods and their sensitivity to physiology and pathology.
Figure 1. Combined MR profiling and optical imaging system for studying water transport processes in tissue specimen
Test system developed for interrogating and validating quantitative diffusion and exchange MR imaging biomarkers via simultaneous real-time NMR and optical microscopy. a) Setup includes the single-sided magnet (NMR MOUSE, Magritek) and the ex vivo tissue chamber (David Ide, Section on Instrumentation, NIMH), modified 3-D technical drawing of the test chamber. b) Drawing of the solenoid radiofrequency (RF) coil containing a viable mouse spinal cord specimen, showing a gap in the RF coil where the sample is imaged. c) Technical drawing of the experimental setup showing the single-sided permanent magnet projecting a magnetic field on the sample from below and the inverted microscope imaging the sample from above. d) Representative sample. e) mean ± SD of AXR (6-pt moving average), Apparent Diffusion Coefficient (ADC), Intrinsic Optical Signal (IOS), and [Ca2+i] fluorophore signal during 20 min 50 mM KCl washin and washout (n = 12, three of which included [Ca2+i] imaging). Insets show zoomed regions with [Ca2+i] inverted for comparison. Arrows in a) mark signal minima and maxima. The data reveal connections between the metrics and their sensitivity to pathological cellular swelling during depolarization. The study helps facilitate the translation from cellular neurophysiology findings to the clinic, and to provide a framework to validate associated MRI methods and their sensitivity to physiology and pathology.
Virtual in vivo MRI histology
Our most mature and widely disseminated in vivo MRI technology is Diffusion Tensor MRI (DTI), with which we measure and map D, a diffusion tensor of water, throughout an imaging volume, often the entire human brain. In neuroradiology, the mean apparent diffusion constant (mADC) is the most widely used diffusion imaging parameter to identify ischemic areas in the brain during acute stroke, to diagnose cancer, and to follow the patient's response to cancer therapy. The mADC is becoming increasingly used to diagnose cancers, such as multiple myeloma, and assess disease progression and response to therapy. Measures of diffusion anisotropy we first proposed (e.g., the fractional anisotropy or FA) are widely used to follow changes in normally and abnormally developing white matter and in many other clinical and neuroscience applications, such as brain white-matter visualization. Our group also pioneered the use of fiber direction–encoded color (DEC) maps to display the orientation of white matter pathways in the brain. To assess anatomical connectivity among various cortical and deep-brain gray-matter areas, We also developed DTI ‘Streamline’ Tractography, which is used to track white-matter pathways and establish ‘anatomical connectivity,' to help neurosurgeons plan brain surgeries, and radiation oncologists to plan radiation dosing, so as to spare ‘eloquent’ parts of the brain. These imaging advances helped inspire several large federally-funded research initiatives, including the NIH Human Connectome Project (HCP) and, more recently, the NIH Brain Initiative.
More recently, we invented and developed a family of advanced in vivo diffusion MRI methods to measure fine-scale microstructural features of axons and other central nervous system (CNS) components (also known as ‘microstructure imaging’), which otherwise could only be assessed using ex vivo histological or pathological methods. These morphological features are generally orders of magnitude smaller than the imaging voxel, but can still be fleshed out using the migration of water molecules to "palpate" them. For example, our Mean Apparent Propagator’ (MAP) MRI is an approach that can detect subtle microstructural and architectural features in both gray and white matter at the micron scale. MAP MRI also subsumes DTI, as well as providing new in vivo quantitative imaging biomarkers to measure and map. We also developed CHARMED MRI, which measures the average axon diameter (AAD), and AxCaliber MRI, which measures the axon-diameter distribution (ADD) along white-matter pathways. The ADD is functionally important, given that axon diameter is a critical determinant of action potential conduction velocity and therefore the rate at which information is transmitted along axon bundles throughout the brain. The ADD also helps determine the latencies or time delays of neural impulses between and among different connected brain areas. These considerations led us to propose a novel MRI–based method to measure the ‘latency connectome,’ including a latency matrix that reports conduction times between different brain areas. We also derived a statistical distribution to explain the form of ADDs observed in different fascicles, suggesting that they broker a trade-off between maximizing information transmission and minimizing metabolic demands.
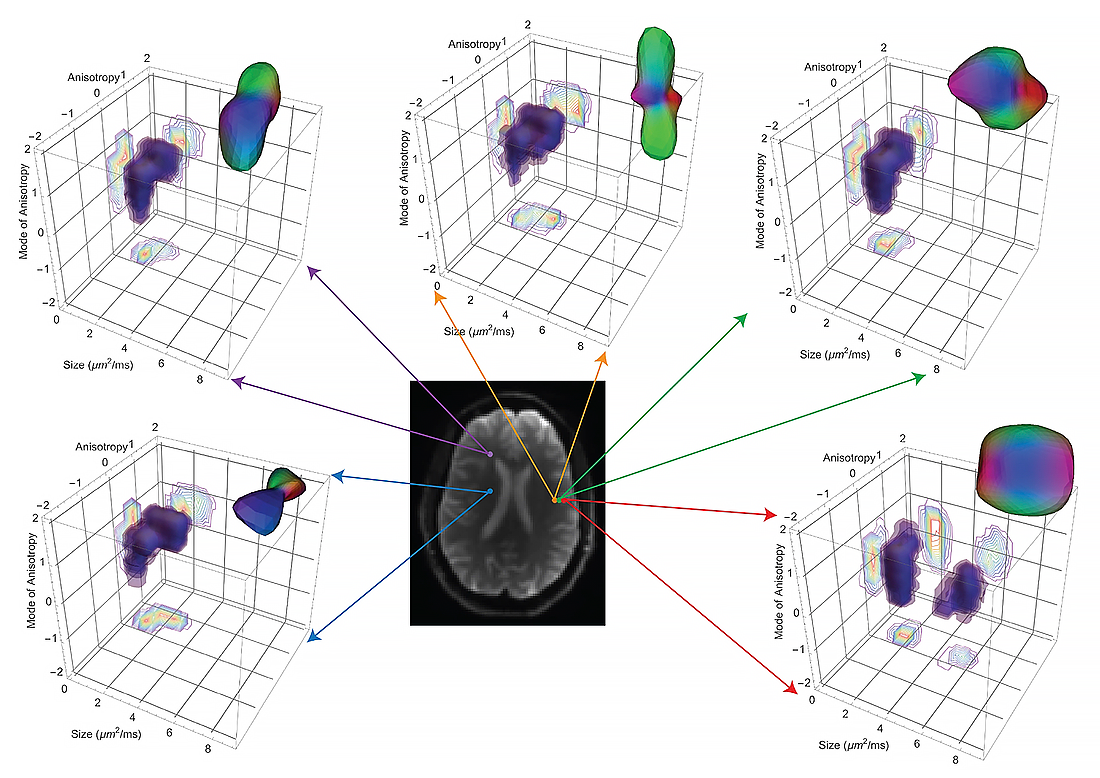
Figure 2. Measuring and mapping the diffusion tensor distribution in the brain in vivo
The Diffusion Tensor Distribution (DTD) measures the mean and covariance structure of diffusion tensors in each voxel within an imaging volume. While we introduced the diffusion tensor distribution (DTD) in the MRI field almost three decades ago, it was only in the past few years that it has been possible to measure this important quantity in vivo owing to the many experimental, conceptual, and technical challenges required to perform this task in a reasonable period of time, and at sufficient accuracy and precision throughout the brain. This reconstruction is obtained in an axial human brain slice whose non-diffusion weighted image is shown at the center. A set of five voxels were chosen in different brain regions corresponding to white matter, gray matter, and CSF. The size-shape distribution of the 6D-DTD is visualized using iso-surfaces or contours in the 3D parameter space of size, anisotropy, mode of anisotropy of the diffusion tensors, and by the micro-orientation distribution function (μODF) computed from ensemble averaging the DTD.
Figure 2. Measuring and mapping the diffusion tensor distribution in the brain in vivo
The Diffusion Tensor Distribution (DTD) measures the mean and covariance structure of diffusion tensors in each voxel within an imaging volume. While we introduced the diffusion tensor distribution (DTD) in the MRI field almost three decades ago, it was only in the past few years that it has been possible to measure this important quantity in vivo owing to the many experimental, conceptual, and technical challenges required to perform this task in a reasonable period of time, and at sufficient accuracy and precision throughout the brain. This reconstruction is obtained in an axial human brain slice whose non-diffusion weighted image is shown at the center. A set of five voxels were chosen in different brain regions corresponding to white matter, gray matter, and CSF. The size-shape distribution of the 6D-DTD is visualized using iso-surfaces or contours in the 3D parameter space of size, anisotropy, mode of anisotropy of the diffusion tensors, and by the micro-orientation distribution function (μODF) computed from ensemble averaging the DTD.
We also developed novel multiple pulsed-field gradient (mPFG) MR methods and demonstrated their feasibility in vivo on conventional clinical MRI scanners as a further means to extract quantitative features in the CNS. Although gray matter appears featureless using DTI, its microstructure and architecture are rich and varied throughout the brain, both along the brain's cortical surface and within the brain's deep subcortical regions. We have been developing several non-invasive, in vivo methods to measure features of cortical gray-matter microstructure and architectural organization that are plainly visible in electron micrographs (EM) but currently invisible in conventional MRI. One example is diffusion tensor distribution (DTD) MRI, in which we use our novel normal tensor-variate distribution to characterize heterogeneity within complex tissue voxels. One of our long-term goals is to segment the cerebral cortex in vivo into its approximately 500 distinct cyto-architectonic areas using such non-invasive imaging methods by developing advanced MRI sequences and analysis pipelines to assess variability in tissue properties in the cortex, such as CORTEC, and other approaches that probe correlations among relaxivities and diffusivities of different water pools. In the latter case, we use multi-dimensional MRI relaxometry and diffusometry to study relaxation processes and water mobility in gray and white matter. We continue to translate these and other methods to the clinic to help identify changes in normal and abnormal development, as well as in inflammation and trauma by developing radiological-pathological correlations between MR and neuropathological images of traumatic brain injury (TBI) tissue specimens. We are in the process of migrating many of these methods both to pre-clinical and clinical applications under the auspices of the Military Traumatic Brain Injury Initiative (MTBI 2) through a collaboration among the Uniformed Services University of Health Sciences (USUHS), the Henry Jackson Foundation (HJF), NICHD, and NIA. Kadharbatcha Saleem and Alexandru Avram have developed imaging tools to correlate high-resolution MAP MRI and histology imaging data, advancing our goal of, one-day, performing virtual in vivo whole-brain histology.
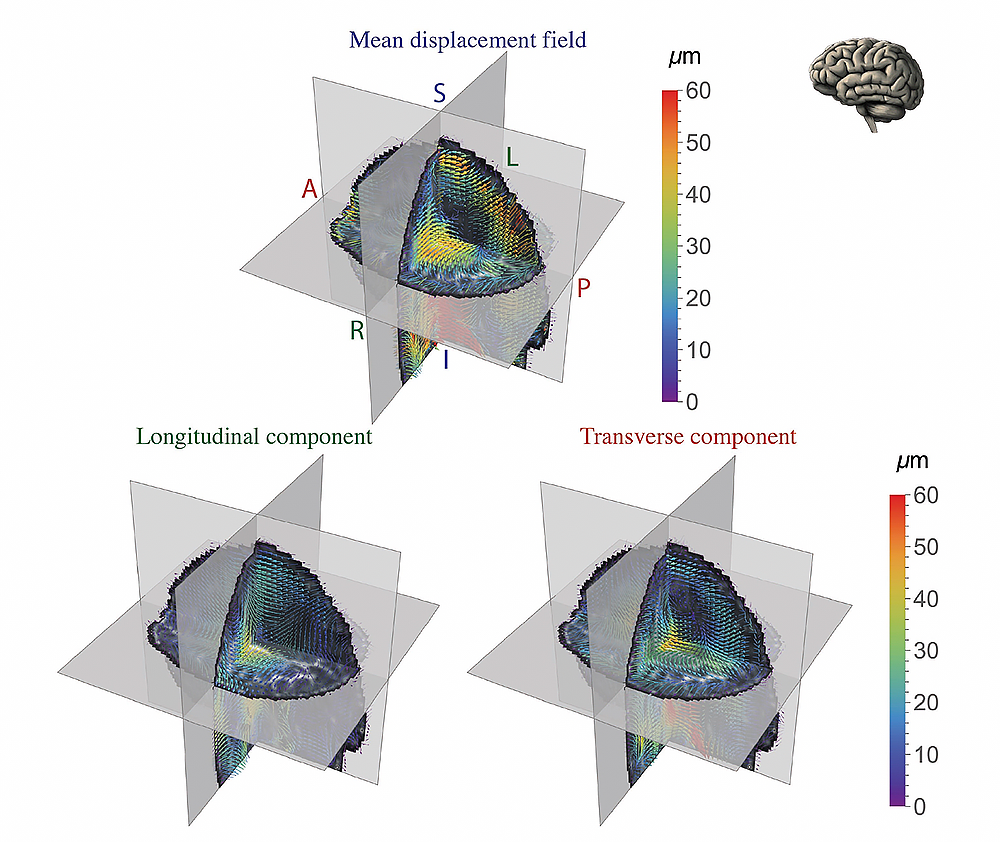
Figure 3. Vector plots showing the 3-D displacement field at a point in the cardiac cycle
We recently combined the ability to measure tissue bulk motion and random motions caused by diffusion. Shown are 3D vector plots of the real part of the mean, longitudinal, and transverse displacement field oscillating at the cardiac frequency (i.e., ≈ 1 Hz) for a representative healthy volunteer. The fractional anisotropy (FA) map from DTI obtained at the quiescent phase of the cardiac cycle is overlaid on the displacement field to provide anatomical context. The displacement vectors are colored based on their magnitude, and anatomical axes (A - Anterior, P - Posterior, S - Superior, I - Inferior, R - Right, L - Left) are indicated. The mean displacement field had a funnel-shaped profile along the I–S axis. The longitudinal component of displacement was non-zero, albeit with smaller magnitude than the transverse component.
Figure 3. Vector plots showing the 3-D displacement field at a point in the cardiac cycle
We recently combined the ability to measure tissue bulk motion and random motions caused by diffusion. Shown are 3D vector plots of the real part of the mean, longitudinal, and transverse displacement field oscillating at the cardiac frequency (i.e., ≈ 1 Hz) for a representative healthy volunteer. The fractional anisotropy (FA) map from DTI obtained at the quiescent phase of the cardiac cycle is overlaid on the displacement field to provide anatomical context. The displacement vectors are colored based on their magnitude, and anatomical axes (A - Anterior, P - Posterior, S - Superior, I - Inferior, R - Right, L - Left) are indicated. The mean displacement field had a funnel-shaped profile along the I–S axis. The longitudinal component of displacement was non-zero, albeit with smaller magnitude than the transverse component.
Quantitative MRI biomarker development
Clinical MRI still lacks the quantitative character of computed tomographic (CT) imaging, revealing only either gross morphological features or focal abnormalities. With ‘quantitative’ clinical MRI techniques, one measures and maps meaningful intrinsic physical quantities or chemical variables that possess physical units and can be compared among different tissue regions. Quantitative MRI methods such as DTI also increase sensitivity, providing a basis for monitoring subtle changes that occur, e.g., during the progression or remission of disease. Quantitative MRI methods should also continue to advance ‘precision imaging’ studies, in which MRI phenotypic and genotypic data can be meaningfully incorporated and used for improved diagnostic and prognostic assessments, such as in the NIH ENIGMA (Enhanced NeuroImaging Genetics through MetaAnalysis) project.
We developed numerical and statistical methods, including algorithms that generate a continuous, smooth approximation to the discrete, noisy, measured DTI field data, so as to reduce noise, and which allowed us to implement ‘Streamline’ Tractography. We proposed a novel Gaussian distribution for tensor-valued random variables that we use to design optimal DTI experiments and interpret their results. We and others recently re-purposed this framework to characterize heterogeneity of diffusion properties within individual voxels. In tandem, we developed non-parametric empirical (e.g., Bootstrap) methods to determine the statistical distribution of DTI–derived quantities in order to study, e.g., the variability and reliability of computed white-matter trajectories to assess the statistical significance of findings in a wide range of biological and clinical applications that had previously been tested using ad hoc statistical methods. We are also developing novel methods to register different brain volumes and to generate group-averaged DTI data or atlases from various subject populations, based on the Kullback-Leibler divergence and other distance metrics, like the “earthmover's distance.”
Previously, we carried out clinical studies that utilize novel quantitative MRI acquisition and analysis methods and whose aim is to improve accuracy and reproducibility of diagnosis and to detect and follow normal and abnormal development in children and adolescents. One early example is the NIH Study of Normal Brain Development, jointly sponsored by the NICHD, NIMH, NINDS, and NIDA, which was initiated in 1998. The Brain Development Cooperative Group (archived June 24, 2024) is still publishing, primarily by mining the MRI data, many of which our lab processed and contributed, serving as the study's DTI Data-Processing Center (DPC). The processed DTI data were uploaded to the National Database for Autism Research (NDAR). Our former Staff Scientist Carlo Pierpaoli, who spearheaded this work, continues to support, update, and disseminate the processing and analysis software called “TORTOISE,” which grew out of our efforts and can be downloaded from https://tortoise.nibib.nih.gov. We continue to improve our ability to follow normal trajectories in pediatric brain development through an active collaboration between the Gates Foundation (GF) and NICHD. Our role in this UNITY project is to develop new MRI pulse sequences and analysis pipelines to probe the state of different water populations in tissue, which has the potential to follow normal and abnormal pediatric brain developmental trajectories.
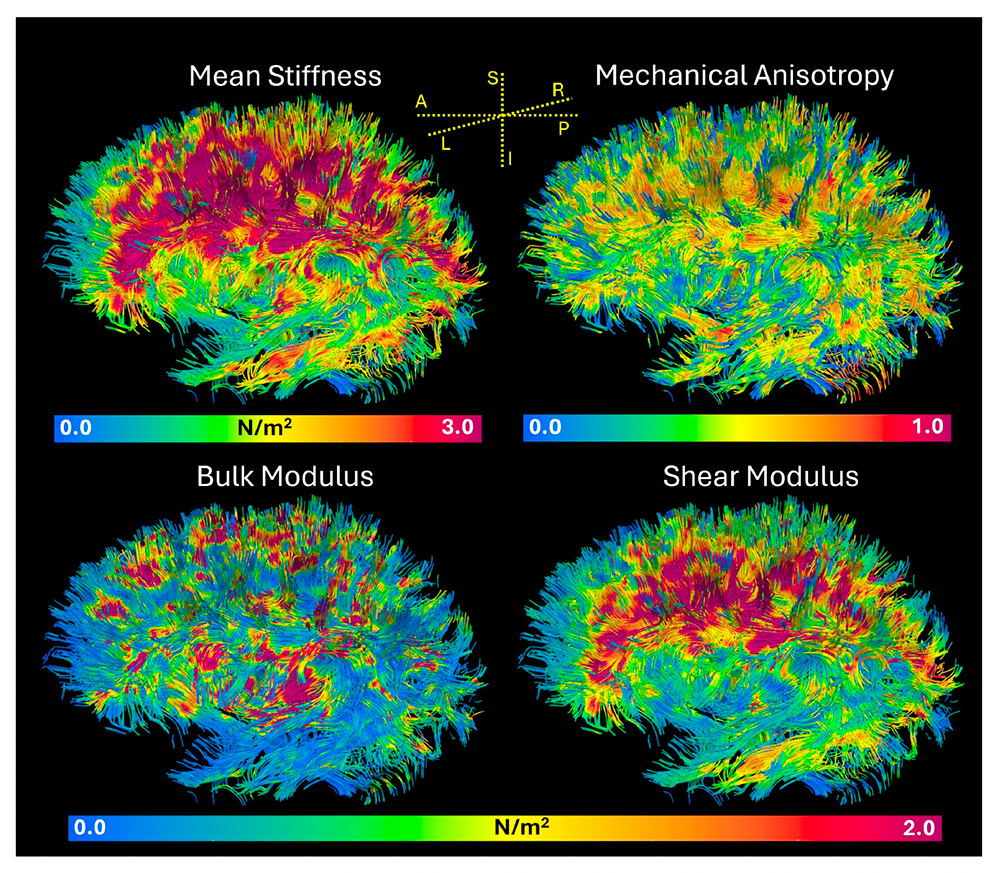
Figure 4. In vivo measured mechanical stiffness of white matter pathways shown using Diffusion Tensor MRI (DTI) tractography
Stiffness of fiber tracts in the human brain visualized by coloring the diffusion tensor tractograms according to the local average stiffness, mechanical anisotropy, bulk, and shear modulus measured using Magnetic Resonance Elastography (MRE). The internal capsule and corona radiata fibers are stiffer than others. The tracts exhibit different degrees of mechanical anisotropy. The bulk and shear modulus were highly heterogeneous across tracts. It is important to note that these measurements are made in vivo using conventional clinical MRI scanners without using an external tamper or vibrator to induce shear waves in the brain.
Figure 4. In vivo measured mechanical stiffness of white matter pathways shown using Diffusion Tensor MRI (DTI) tractography
Stiffness of fiber tracts in the human brain visualized by coloring the diffusion tensor tractograms according to the local average stiffness, mechanical anisotropy, bulk, and shear modulus measured using Magnetic Resonance Elastography (MRE). The internal capsule and corona radiata fibers are stiffer than others. The tracts exhibit different degrees of mechanical anisotropy. The bulk and shear modulus were highly heterogeneous across tracts. It is important to note that these measurements are made in vivo using conventional clinical MRI scanners without using an external tamper or vibrator to induce shear waves in the brain.
Traumatic Brain Injury (TBI) represents a significant public health challenge for our pediatric population, but also for those serving in the military. Our TBI research, particularly in detecting mild TBI (mTBI), has continued to expand through partnerships with various Department of Defense (DoD) entities and the GE-NFL–mTBI (General Electric–National Football League Head Health Initiative) MRI project. Diffusion MRI (dMRI) provides essential information to aid in the assessment of TBI, but conventional dMRI methods have lacked sufficient specificity. To improve the accuracy and reproducibility of dMRI we developed a data-processing pipeline, and, in collaboration with the DoW Military Traumatic Brain Injury Initiative (MTBI 2), we helped incorporate MAP-MRI into our NICHD TORTOISE pipeline to detect sequelae of mTBI. We have also been employing multi-dimensional MRI relaxometry-diffusometry methods to study the etiology of various types of TBI, in collaboration with the USUHS Neuropathology Research Division and under the auspices of MTBI 2, and to improve the correlation and integration of neuropathology and neuro-radiological imaging data. We also partnered with MTBI 2 to study ways to measure very slow flows that occur during glymphatic transport, a mechanism that the brain uses to wash away harmful macromolecules and cellular components. With our partners at the University of Arizona, this pre-clinical research is providing experimental data to enable us to migrate these imaging approaches to the clinic.
In a long-term collaboration with Sara Inati, who studies focal epilepsy, a devastating disorder that is difficult to detect using conventional neuroradiological methods, we are developing and testing various new MRI–based methods that we believe may reveal pathological changes in brain microstructure and architectural organization in this disorder, for example in cortical dysplasia, to improve localization and assessment of cortical lesions. We are currently testing the utility of DTD MRI approach on Sara Inati's patient cohort.
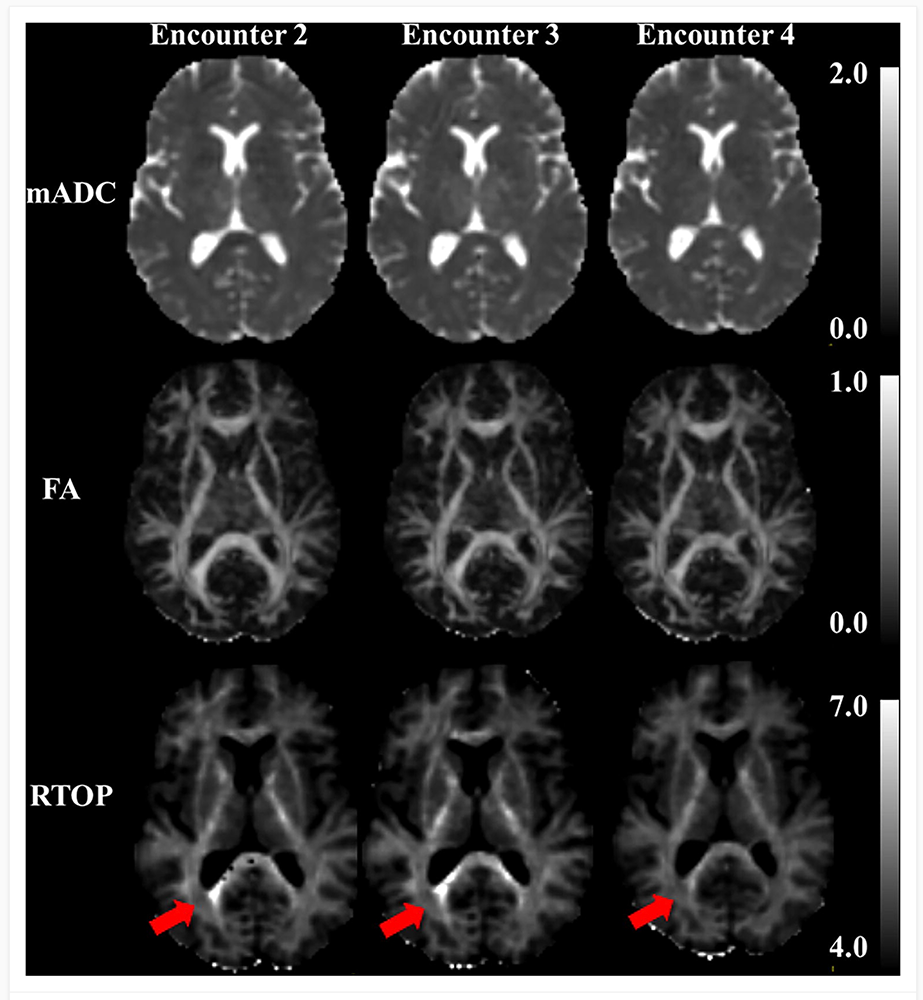
Figure 5. Trying to make the invisible visible: imaging mild traumatic brain injury
Scans of subjects with mild traumatic brain injury obtained during several imaging sessions or episodes. Mean Apparent Propagator MRI is used to provide quantitative imaging biomarkers hypothesized to be able to follow microstructural changes occurring in this condition. The mean apparent diffusivity (mAD) and fractional anisotropy (FA) from diffusion tensor MRI (DTI) often do not show evidence of microstructure changes associated with injury in mTBI. The more sensitive and specific quantitative imaging biomarkers, such as the return to origin probability (RTOP), may provide additional information with which to make such diagnoses.
Figure 5. Trying to make the invisible visible: imaging mild traumatic brain injury
Scans of subjects with mild traumatic brain injury obtained during several imaging sessions or episodes. Mean Apparent Propagator MRI is used to provide quantitative imaging biomarkers hypothesized to be able to follow microstructural changes occurring in this condition. The mean apparent diffusivity (mAD) and fractional anisotropy (FA) from diffusion tensor MRI (DTI) often do not show evidence of microstructure changes associated with injury in mTBI. The more sensitive and specific quantitative imaging biomarkers, such as the return to origin probability (RTOP), may provide additional information with which to make such diagnoses.
Other quantitative biomarkers arise from characterizing different material or mechanical properties of brain tissue in vivo. Magdoom Kulam developed a novel MRI pipeline, magnetic resonance elastography (MRE), to measure the distribution of tissue stiffness throughout the brain without the use of a mechanical actuator, a method that we tested on polymeric and tissue phantoms, and clinical tests were performed. Recently, this approach was implemented on clinical 3T and 7T MRI scanners to measure and map material properties of the brains of normal human subjects.
Biopolymer physics: water-ion-biopolymer interactions
Despite their crucial role in biological processes, little is still known about the physics of water-ion-biopolymer interactions in the physiological ionic-strength regime. To determine the effect of ions on the structure and dynamics of key biopolymers, we combined macroscopic techniques (osmotic swelling-pressure and mechanical measurements) with high-resolution scattering methods (e.g., SANS and SAXS, SLS, and DLS). Macroscopic swelling-pressure measurements provide information about the system's overall thermodynamic response, while SANS, SAXS, SLS, and DLS probe biopolymer structure at molecular and atomic through supramolecular length scales to quantify the effect of environmental changes (e.g., in ion concentration, ion valance, pH, temperature) on the interactions among biopolymers, water, and ions.
These basic studies on bio-mimetic tissue analogs have led to the development of novel MRI phantoms designed to support our quantitative imaging program, including diffusion MRI phantoms, which we use to calibrate MRI scanners to assure the quality and fidelity of their imaging data in single-subject, longitudinal, and multi-site studies. For instance, our U.S. Patent for a ‘Phantom for Diffusion MRI Imaging’ is now incorporated in CaliberMRI phantoms, enabling quantitative diffusion MRI studies at many clinical sites worldwide. Our colleagues at NIST Boulder have also incorporated our polyvinylpyrrolidone (PVP) polymer into their own diffusion MRI NIST "standard". We have used various glass microcapillary array (GCA) geometries to mimic features of axons in white-matter pathways in order to interrogate our AxCaliber and dPFG MRI models of water diffusion in such white matter pathways, and, more recently, 3-D printed polymeric phantoms, and multi-component polymer gel systems exhibiting inter compartmental water exchange to calibrate our novel Diffusion Exchange Spectroscopy (DEXSY) MRI acquisitions.
Measuring and mapping functional properties of extracellular matrix
We study interactions among extracellular matrix (ECM) components, having initially used cartilage as a model system because it is aneural, avascular, and nearly acellular. In cartilage ECM, collagen (type II) is organized into fiber bundles that form a network that entraps the major proteoglycan (PG), the bottlebrush-shaped aggrecan molecule. The biomechanical behavior of cartilage and other ECMs reflects their molecular composition and microstructure, which change during development, disease, degeneration, and aging. To determine tissue structure/function relationships, we measured physical/chemical properties of ECM tissues and tissue analogs at different length- and time-scales, using, e.g., osmometry, SANS, SAXS, neutron spin-echo (NSE), SLS, DLS, and AFM. Understanding the physical and chemical mechanisms affecting cartilage swelling (hydration) is essential to predicting its load-bearing and lubricating ability, which are mainly governed by osmotic and electrostatic forces. To quantify the effect of hydration on cartilage properties, we previously developed a novel tissue micro-osmometer to measure the effect of equilibrium water activity (vapor pressure) in small tissue samples (less than 1 µg). We also make osmotic-pressure measurements to determine how the individual components of cartilage ECM (e.g., aggrecan and collagen) contribute to the total load-bearing capacity of the tissue. We also demonstrated that aggrecan-hyaluronic aggregates self-assemble into microgel particles, contributing to improved dimensional stability and to the lubricating ability of cartilage. We found that aggrecan is highly insensitive to changes in the ionic environment, particularly to divalent cations such as calcium, which is critical for maintaining the tissue's mechanical integrity, and allows aggrecan to serve as a calcium-ion reservoir in cartilage and bone.
More recently, to model cartilage ECM, Ferenc Horkay invented and developed a new biomimetic composite material consisting of polyacrylic acid (PAA) micro-gel particles dispersed and embedded within a polyvinyl alcohol (PVA) fibrous gel network. PAA mimics cartilage's proteoglycan phase (i.e., hyaluronic acid–aggrecan complexes), while PVA mimics the fibrous collagen network phase entrapping them. The PVA/PAA biomimetic composite model system reproduces both the shape of the cartilage-swelling pressure curves and the stiffness values reported for healthy and osteoarthritic human cartilage specimens. Studies on these model composite hydrogels yield invaluable insights into how macromolecular factors (matrix stiffness, swelling pressure, fixed-charge density, etc.) can affect the tissue's macroscopic mechanical/swelling properties, and ultimately its load-bearing and lubricating abilities, and their loss in various diseases and disorders, including osteoarthritis.
We are now attempting to develop and design novel non-invasive MRI methods, with the aim of inferring parameters characterizing ECM composition, patency, and functional properties in vivo. Our goal is to use MRI for early diagnosis of ECM diseases and to follow normal and abnormal ECM development, which entails making components of ECM (e.g., collagen and PGs) that are ‘invisible’ to MR ‘visible’ so as to predict the functional properties of the composite tissue, such as its load-bearing ability. An obstacle is that protons bound to immobile species (e.g., collagen) are largely invisible with conventional MRI methods. However, magnetization exchange (MEX) MRI (as well as other related methods) now make it possible to detect bound protons indirectly by transferring their magnetization to the abundant free water protons surrounding them, which enables us to quantitate collagen content in tissue. In previous pilot studies with Uzi Eliav (deceased) and Ed Mertz, we applied the new MEX MRI method to determine the concentration and distribution of the main macromolecular constituents in bovine femoral-head cartilage samples. The results were qualitatively consistent with those obtained by histological techniques, such as high-definition infrared (HDIRI) spectroscopic imaging. Our approach has the potential to map tissue structure and functional properties in vivo and non-invasively. Recent molecular dynamics (MD)–based models of cartilage and cartilage ECM analogs have helped us interpret our experimental findings in terms of molecular interactions and processes. Currently, Julian Rey is using Finite Element Models (FEM) to describe the behavior of these novel composite materials to explain and predict their properties.
We have also been employing several novel MR–based methodologies to characterize ECM properties using our one-sided NMR profiling systems to study water relaxation, diffusion, and exchange processes in various tissue specimens. We are adapting these methods to study the composition and organization of brain ECM, which is a complex medium about which surprisingly little is known. There is even disagreement about the composition and molecular architecture of brain ECM. We hope to make this critically important invisible component visible using various novel MRI methods now under development.
Patents
- Time efficient multi-pulsed field gradient (mPFG) MRI without concomitant gradient field artifacts
MMK Najmudeen, PJ Basser, ME Komlosh, US Patent 12,493,092, 2025 - Multidimensional MRI signature for specific detection of traumatic brain injury in vivo
DH Benjamini, PJ Basser, D Iacono, US Patent 12,343,132 - MRI tractography based transit time determination for nerve fibers
PJ Basser, US Patent 12,345,788 - Nuclear magnetic resonance methods of determining homeostatic perturbations
R Ravin, NH Williamson, TX Cai, PJ Basser, US Patent App. 18/697,168
Additional Funding
- “Connectome 2.0: Developing the next generation human MRI scanner for bridging studies of the micro-, meso- and macro-connectome.” NIH BRAIN Initiative-funded 1U01EB026996-01
- “Advanced Neuroimaging Research and Development Core” MTBI2/USUHS/HJF/DoD
Publications
- Effect of divalent ions on the structure of polyelectrolyte gels. Polymer 2025 327:128311
- Passive water exchange between multiple sites can explain why apparent exchange rate constants depend on ionic and osmotic conditions in gray matter. Magn Reson Lett 2025 200225;in press
- In vivo palpation of anisotropic human brain tissue using MRI. bioRxiv 2025 doi: 10.1101/2025.06.20.660588;preprint
- Measuring the velocity autocorrelation function using diffusion NMR. J Chem Phys 2025 162(17):174203
- Modeling osmotic deswelling of composite hydrogels with finite element analysis. Polymer (Guildf) 2025 333:128634
Collaborators
- Dan Benjamini, PhD, Unit on Multiscale Imaging and Integrative Biophysics, NIA, Bethesda, MD
- Jack Douglas, PhD, Materials Science and Engineering Division, NIST, Gaithersburg, MD
- Brian Edlow, MD, Massachusetts General Hospital, Boston, MA
- Dario Gasbarra, PhD, University of Helsinki, Helsinki, Finland
- David Feinberg, PhD, University of California, Berkeley, CA
- Silvina Horowitz, PhD, Mobile Clinical Research Unit, NINDS, Bethesda, MD
- Susie Huang, MD PhD, Martinos Center, Harvard Medical School, Boston, MA
- Beth Hutchinson, PhD, University of Arizona, Tucson, AZ
- Sara Inati, MD, Electroencephalography (EEG) Section, NINDS, Bethesda, MD
- Mustapha Irfanoglu, PhD, Laboratory on Quantitative Medical Imaging, NIBIB, Bethesda, MD
- Edward Mertz, PhD, Section on Physical Biochemistry, NICHD, Bethesda, MD
- Nicole Y. Morgan, PhD, Trans-NIH Shared Resource on Biomedical Engineering and Physical Science (BEPS), NIBIB, Bethesda, MD
- Evren Özarslan, PhD, Linköping University, Linköping, Sweden
- Sinisa Pajevic, PhD, Section on Critical Brain Dynamics, NIMH, Bethesda, MD
- Daniel Perl, MD, Uniformed Services University of the Health Sciences, Bethesda, MD
- Carlo Pierpaoli, MD, PhD, Laboratory on Quantitative Medical Imaging, NIBIB, Bethesda, MD
- Dietmar Plenz, PhD, Section on Critical Brain Dynamics, NIMH, Bethesda, MD
- Randall Pursley, MS, Instrumentation Development and Engineering Application Solutions (IDEAS), NIBIB, Bethesda, MD
- Joelle Sarlls, PhD, In Vivo NMR Center, NINDS, Bethesda, MD
- Brain Development Cooperative Group, Various
Contact
For more information, email basserp@mail.nih.gov or visit https://www.nichd.nih.gov/research/atNICHD/Investigators/basser.