Neuronal Circuits Controlling Behavior: Genetic Analysis in Zebrafish

- Harold Burgess,
PhD, Head, Section on Behavioral Neurogenetics - Tripti Gupta, PhD, Staff Scientist
- Sreerag Othayoth Vasu, PhD, Visiting Fellow
- Madalina Robea, PhD, Visiting Fellow
- Svetlana Semenova, PhD, Visiting Fellow
- Hariom Sharma, PhD, Visiting Fellow
- Reid Doctor, BSc, Graduate Student
- Giovannina Kirby, BSc, Postbaccalaureate Fellow
- Jennifer L. Sinclair, MSc, Zebrafish Technician
The Section on Behavioral Neurogenetics studies the development and function of neural circuits for sensory processing, and how these are modified in neurodevelopmental disorders. We use the larval zebrafish as a model because its brain exhibits the basic architecture of the vertebrate brain but is much less complex than the mammalian brain. Despite the relative simplicity of their nervous system, zebrafish have a sophisticated repertoire of sensory-guided and internally driven behaviors. Furthermore, the optical clarity of the embryo facilitates visualization of individual neurons and their manipulation with genetic techniques. Behavior in larvae is innate and thus exhibits minimal variability between fish. Subtle alterations in behavior can therefore be robustly measured, making it possible to quickly assess the contribution of identified neurons to a variety of motor behaviors.
We focus on two aspects of behavioral regulation: the neuronal mechanisms by which sensory context regulates behavioral decisions; and the pathways that sustain changes in behavioral state. Neuronal connections that allow the brain to integrate sensory and internal-state information are established through genetic interactions during development and are frequently disrupted by gene mutations associated with neurodevelopmental disorders. We can therefore use discoveries about sensory motor integration pathways to understand how human disease genes disrupt brain development, and we frequently collaborate with clinical investigators to generate and study zebrafish to provide new insights into human disease genes.
To support these objectives, we develop new genetic tools and behavioral assays to probe the nexus between neuronal function and behavior. We are especially interested in technologies that enable us to probe circuit and gene function at single-cell resolution, using transgenic expression of effectors as well as classical genetic methods to achieve this.
Zebrafish models of neurodevelopmental disorders
We collaborate widely with clinicians to generate and characterize zebrafish models for mutations discovered in humans (often through exome sequencing) that are likely to have a neurodevelopmental origin. We use the CRISPR/Cas9 system to generate lesions in zebrafish genes that are homologous to those disrupted in the human disorders. We then apply behavioral analysis, transcriptomics, and voxel-based morphometry as part of a broad phenotyping strategy. A unique feature of brain imaging in zebrafish is the ability to visualize the total architecture of the brain while simultaneously recording the position and morphology of every constituent labeled neuron. Such high precision of alignment permits statistically robust whole-brain analysis of neuronal composition and morphology in zebrafish mutant models, pinpointing brain regions with changes that are difficult to detect visually. The technique can be applied to almost any zebrafish neurodevelopmental model, thereby enabling robust and quantitative detection of subtle changes in brain structure or composition. Through this work, we aim to provide insight into the fundamental molecular and cellular processes associated with each disorder.
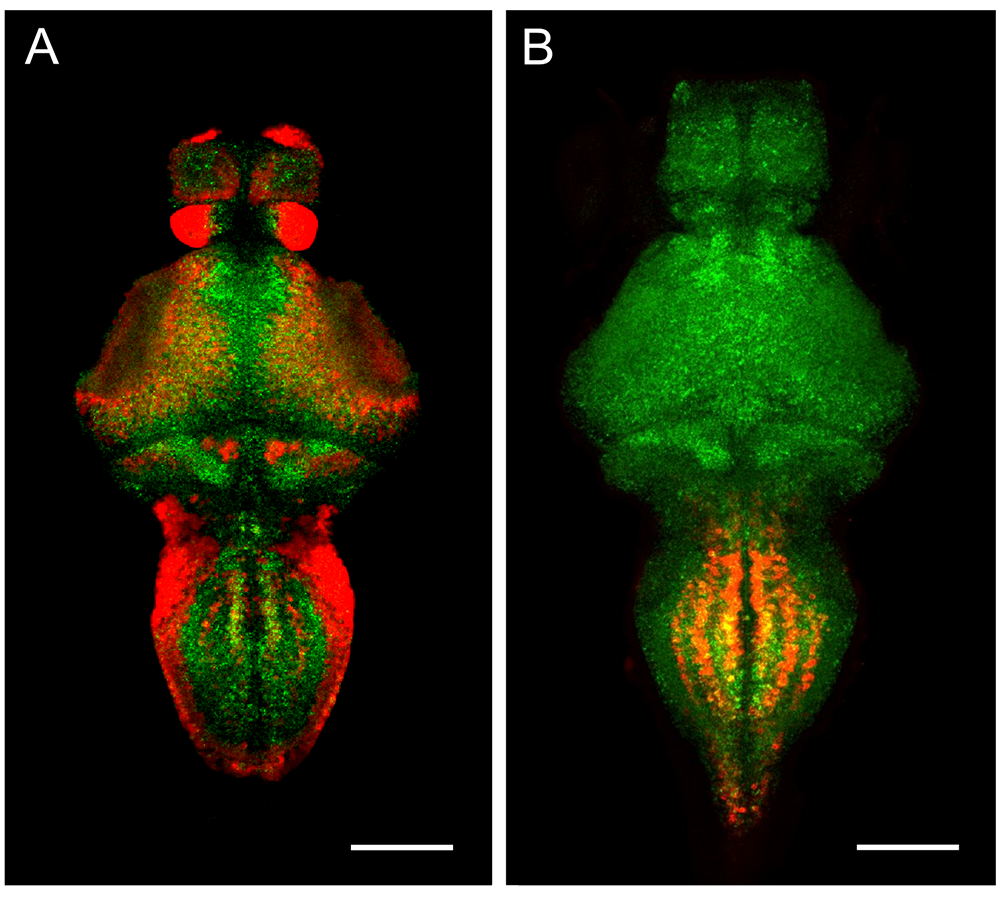
Using voxelwise morphometry, we discovered that mutations in xrcc1, the zebrafish equivalent of a human disease gene, leads to a highly localized reduction in tissue volume in the larval cerebellum. Xrcc1 is a scaffold protein that negatively regulates Parp1, a protein involved in single-strand break repair. We showed that cerebellar defects in xrcc1 mutants can be rescued by simultaneously knocking down parp1, suggesting that PARP1 inhibitors might be useful therapeutic candidates for XRCC1–related disorders [Reference 1]. We also studied srrm4, which earlier studies had suggested encodes a key regulator of alternatively spliced microexons during neural development that were implicated in the etiology of autism. We found that srrm4 is indeed expressed during neural development in zebrafish, enriched in granule populations in the cerebellum and torus longitudinalis (Figure 1). Previous studies had uncovered roles for srrm4 in zebrafish by interfering with gene function using microinjection of antisense ‘morpholino’ oligonucleotides. However, these phenotypes failed to manifest in larvae with srrm4 gene mutations, and indeed, we found that loss of srrm4 had little effect on neural development or micro-exon splicing in zebrafish [Reference 2].
Figure 1. srrm4 partially co-localizes with glutamatergic and glycinergic neuron markers in the medulla.
Dorsal views of single confocal planes from (A) srrm4 and vglut2a:GFP expression, and (B) srrm4 and glyt2:RFP expression, revealing widespread expression of the microexon splicing factor across a range of different cell types.
Figure 1. srrm4 partially co-localizes with glutamatergic and glycinergic neuron markers in the medulla.
Dorsal views of single confocal planes from (A) srrm4 and vglut2a:GFP expression, and (B) srrm4 and glyt2:RFP expression, revealing widespread expression of the microexon splicing factor across a range of different cell types.
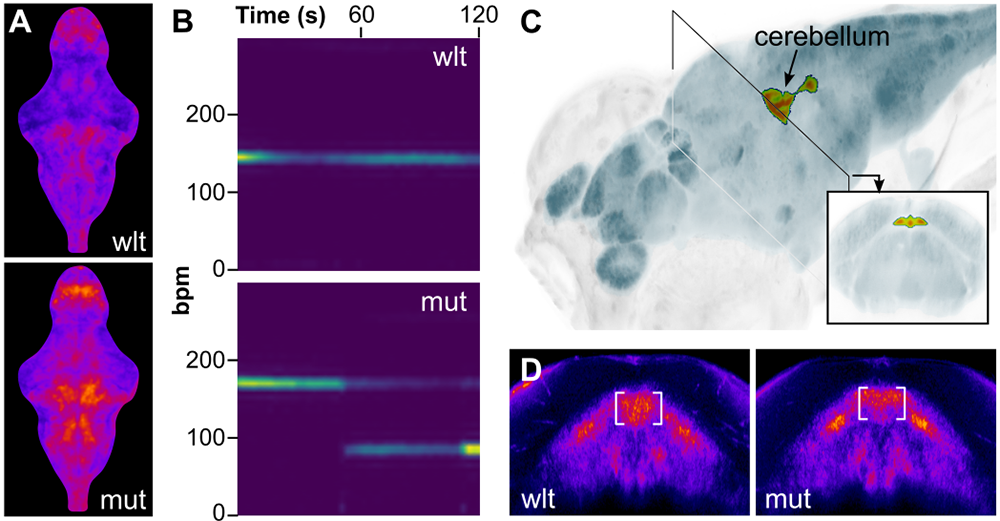
We also characterized a new zebrafish model of Timothy syndrome, a severe multisystem disorder that is caused by a gain-of-function mutation in the pore-forming subunit of the CaV1.2 channel CACNA1C, that prolongs channel opening. The zebrafish line carries a point mutation that precisely recapitulates the mutation found in patients, allowing us to assess physiological and neuromorphometric phenotypes. Consistent with clinical manifestations, larvae carrying the mutation showed heightened neural activity, increased susceptibility to seizure-like behavior and cardiac arrhythmias (Figure 2). Multiple abnormalities were apparent in brain architecture, including cerebellar hypotrophy and abnormal GABA neuron development. Because CaV1.2 channels are known to open spontaneously at elevated temperature, we speculated that fish with the mutation might be especially sensitive to fever-like conditions. Indeed, we found heightened seizure susceptibility and arrhythmia when they were tested at increased temperature, drawing attention to the possibility that fever might be a particular risk factor for Timothy syndrome patients [Reference 3].
Figure 2. A zebrafish knock-in model of Timothy syndrome (TS)
A. Mean change in neuronal activity (ΔF/F, GCaMP imaging) over 5 min. in wild-type and mutant larvae. B. Spectrogram of ventricle contraction frequency (beats per min.) in a wild-type larva and TS mutant with intermittent arrhythmia. C. Decreased cerebellar volume in TS mutant revealed by brain morphometry. D. Reduced en1b expression (HCR in situ, pseudocolored for intensity) at the midbrain-hindbrain boundary in TS mutant.
Figure 2. A zebrafish knock-in model of Timothy syndrome (TS)
A. Mean change in neuronal activity (ΔF/F, GCaMP imaging) over 5 min. in wild-type and mutant larvae. B. Spectrogram of ventricle contraction frequency (beats per min.) in a wild-type larva and TS mutant with intermittent arrhythmia. C. Decreased cerebellar volume in TS mutant revealed by brain morphometry. D. Reduced en1b expression (HCR in situ, pseudocolored for intensity) at the midbrain-hindbrain boundary in TS mutant.
Neuronal pathways for auditory sensory processing
Startle responses are rapid reflexes that are triggered by sudden sensory stimuli, and which help animals defend against, or escape from, potentially threatening stimuli. In both fish and mammals, startle responses are initiated by giant reticulospinal neurons in the medulla, which receive short-latency sensory input from diverse sensory modalities. Although highly stereotyped, startle responses are nevertheless modulated by sensory context and behavioral state and are therefore an excellent system in which to study how such information is integrated. Moreover, the underlying sensorimotor circuits are relatively simple, facilitating studies into the effects of gene mutations that cause neurodevelopmental disorders such as autism and schizophrenia.
In mammals, including humans, the startle response to a strong auditory stimulus can be inhibited by pre-exposure to a weak acoustic ‘prepulse,’ a form of startle modulation termed prepulse inhibition, which is diminished in several neurological conditions. Previously, we showed that, in zebrafish as in mammals, several distinct cellular mechanisms mediate prepulse inhibition, depending on the time interval between the prepulse and the startle stimulus, with NMDA receptor signaling playing a key role for intervals greater than 100 ms. Accordingly, zebrafish with gene mutations in specific NMDA receptor subunits linked to schizophrenia show diminished prepulse inhibition. We have now also identified neurons that mediate prepulse inhibition at intervals of less than 100 ms and confirmed previous reports that a GABAergic mechanism is involved. We demonstrated that these neurons directly connect to central startle command neurons and defined the molecular signaling pathway between these neurons that is required for prepulse inhibition.
Our understanding of the cellular pathway that links auditory sensory cues to startle behavior has important gaps. We therefore defined afferent connectivity from the auditory sensory ganglion in zebrafish (the statoacoustic ganglion), in order to elucidate pathways that transmit behaviorally relevant vibration and acceleration signals to central regions that process sensory information for startle behavior, balance, and eye movements. By using transgenic lines that selectively label subsets of neurons in the statoacoustic ganglion, as well as with targeted laser ablation, we identified specific neurons that transmit abrupt auditory signals to the brain, to initiate escape behavior. Given the small size of the larvae, a challenge is to link neuronal activity to behavior. Thus, to non-invasively record neuronal activity in these experiments, we built a new microscope that allows us to record changes in transgenically expressed GCaMP fluorescence in freely swimming larvae.
New genetic tools for dissecting neural circuit function
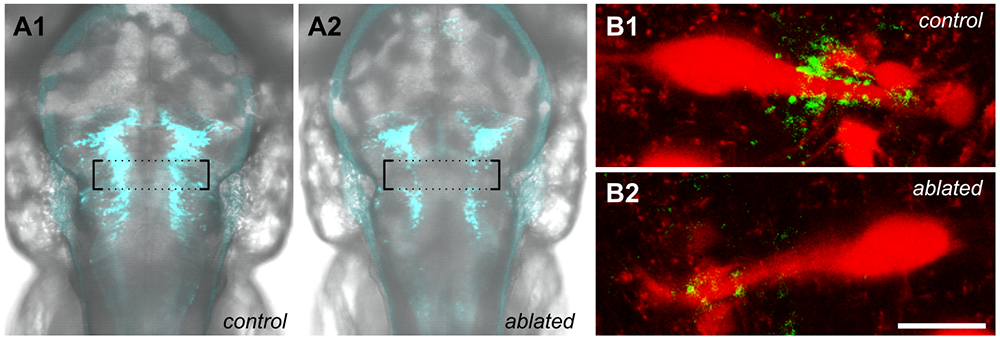
Understanding how the brain utilizes sensory information to guide behavior requires dissecting the precise underlying chain of neuronal signaling. A difficulty is that many neurons are connected to a large number of downstream targets, often making it difficult to conclusively demonstrate that a specific connection is part of a behaviorally relevant pathway. To address this, we developed a new method to selectively eliminate specific contacts between neurons. The techniques utilizes the cell-type–specific expression of the fluorogen-activating protein dL5**. After exposing larvae to a cell-permeable photosensitizer that binds to dL5**, we can use spatially patterned illumination to optically ablate cells or cellular domains. We first demonstrated that cytoplasmic neuronal expression of dL5** enables us to use photoablation to selectively lesion neurons within a larger transgenic pattern, without damaging adjacent dL5** non-expressing cells (Figure 3A). Next, we fused dL5** to synaptophysin to ensure presynaptic localization, aiming to photo-ablate selected output synapses of neurons of interest. We validated the method by eliminating specific output synapses of gsx1–expressing neurons that mediate distinct effects on escape behavior, demonstrating that we could suppress prepulse inhibition without disrupting other signaling events that depend on these neurons (Figure 3B). This is a powerful method and should be applicable to a wide range of questions concerning the transmission of information between neurons across different experimental systems and fields of study [Reference 4].
Figure 3. Spatially localized cell and synapse ablation by transgenic expression of the MG-2I binding protein dL5
A. Rhombomere specific ablation in y252-Gal4, UAS:dL5-mCer control (A1) and light-exposed larva (A2), using a digital micromirror device to restrict illumination to only the indicated area (boxed). B. Untreated y252-Gal4, UAS:syp-dL5-mNeonGreen right Mauthner cell (B1) and left Mauthner cell after photoablation of synapses on lateral dendrite (B2). Left and right cells are from the same larva. Scale bar 20 µm.
Figure 3. Spatially localized cell and synapse ablation by transgenic expression of the MG-2I binding protein dL5
A. Rhombomere specific ablation in y252-Gal4, UAS:dL5-mCer control (A1) and light-exposed larva (A2), using a digital micromirror device to restrict illumination to only the indicated area (boxed). B. Untreated y252-Gal4, UAS:syp-dL5-mNeonGreen right Mauthner cell (B1) and left Mauthner cell after photoablation of synapses on lateral dendrite (B2). Left and right cells are from the same larva. Scale bar 20 µm.
Publications
- Parp1 deletion rescues cerebellar hypotrophy in xrcc1 mutant zebrafish. Sci Rep 2025 15:17043
- Mutations in the microexon splicing regulator srrm4 have minor phenotypic effects on zebrafish neural development. G3 (Bethesda) 2025 15:jkaf052
- Elevated body temperature exacerbates arrhythmia and seizure-like activity in a zebrafish model of Timothy syndrome. bioRxiv 2025 doi: 10.1101/2025.03.11.642683;preprint
- Functional interrogation of neuronal connections by chemoptogenetic presynaptic ablation. bioRxiv 2025 doi: 10.1101/2025.04.04.647277;preprint
Collaborators
- Carsten Bönnemann, MD, Neuromuscular and Neurogenetic Disorders of Childhood Section, NINDS, Bethesda, MD
- Keith Caldecott, PhD, University of Sussex, Brighton, United Kingdom
- Thomas E. Dever, PhD, Section on Protein Biosynthesis, NICHD, Bethesda, MD
- Andy Golden, PhD, Laboratory of Biochemistry and Genetics, NIDDK, Bethesda, MD
- Todd S. Macfarlan, PhD, Section on Mammalian Epigenome Reprogramming, NICHD, Bethesda, MD
- Reza Maroofian, PhD, University College London, London, United Kingdom
- Anne O'Donnell Luria, MD, PhD, Mendelian Genomics Research Center, Broad Institute of MIT, and Harvard University, Cambridge, MA
Contact
For more information, email haroldburgess@mail.nih.gov or visit https://www.nichd.nih.gov/research/atNICHD/Investigators/burgess.