The Role of Metabolism in the Regulation of Oocyte Development and Growth

- Mary Lilly,
PhD, Head, Section on Gamete Development - Chun-Yuan Ting, PhD, Staff Scientist
- Kuikwon Kim, MS, Biologist
- Shu Yang, PhD, Postdoctoral Fellow
- Richard Garcia, BS, Postbaccalaureate Fellow
- Sophie Menendez, BS, Postbaccalaureate Fellow
- Natalie Rowland, BS, Postbaccalaureate Fellow
Our long-term goal is to obtain a comprehensive understanding of how metabolic signaling pathways influence oocyte growth, development, and quality. Chromosome mis-segregation during female meiosis is the leading cause of miscarriages and birth defects in humans. Recent evidence suggests that many meiotic errors occur downstream of defects in oocyte growth and/or in the hormonal signaling pathways that drive differentiation of the oocyte. Thus, understanding how oocyte metabolism and growth impact meiotic progression is essential to studies in both reproductive biology and medicine.
Metabolic dysregulation, including obesity and diabetes, negatively impact human fertility. We use the genetically tractable model organism Drosophila melanogaster to examine how meiotic progression and oocyte development are instructed by the metabolic program of the egg. In mammals, studies on the early stages of oogenesis face serious technical challenges in that entry into the meiotic cycle, meiotic recombination, and the initiation of the highly conserved prophase I arrest all occur during embryogenesis. By contrast, in Drosophila these critical, highly conserved events of early oogenesis all take place continuously within the adult female. Easy access to the early stages of oogenesis, coupled with available genetic and molecular-genetic tools, makes Drosophila an excellent model for studies on the role of metabolism in oocyte development and maintenance.
Additionally, we also use the Drosophila ovary as a model to investigate the tissue-specific regulation of metabolism. Our focus is on gaining a molecular understanding of how the GTPase–activating GATOR complex modulates TORC1 (protein kinase complex that controls protein synthesis) activity and regulates the autophagic-lysosomal pathway.
A toolbox to dissect the tissue-specific regulation of metabolism
In the United States, metabolic dysregulation is a major contributor to morbidity and mortality. Despite its significance, how cellular metabolism is regulated within a complex multicellular organism remains poorly understood. To address this gap, we aim to develop reagents that enable the in vivo investigation of metabolic pathways, in specific cell types, with a particular focus on the ovary and the critical role of metabolism in maintaining oocyte quality. Therefore, over the past year we generated a molecular-genetic resource that provides a versatile platform to dissect TORC1 regulation and its upstream control by the GATOR–Rag GTPase network in distinct metazoan tissues. By genomically tagging pathway components at their endogenous loci, we created reagents that allow visualization of the subcellular dynamics of these proteins under native expression levels. This approach avoids artifacts associated with overexpression and enables cell-type–specific analyses using the GAL4/UAS system (genetic tool that enables ectopic expression of genes) in Drosophila. The addition of the split–GFP system further enhances these capabilities, permitting the real-time visualization of endogenously tagged proteins in specific tissues and developmental contexts. These reagents now make it possible to follow the activity and localization of GATOR2, GATOR1, and the Rag GTPase with high spatial and temporal resolution across diverse cell types, establishing a toolkit that will facilitate mechanistic and physiological studies of TORC1 regulation in vivo. As proof of principle, we have used these reagents to demonstrate that many of the regulatory principles that govern the GATOR-Rag GTPase signaling pathways in mammals are conserved in Drosophila. We are currently using these tools to explore the molecular mechanism by which GATOR2 inhibits GATOR1 and thus functions to activate TORC1 in order to preserve metabolic homeostasis in the ovary.
The GATOR complex: integrating developmental and metabolic signals in oogenesis
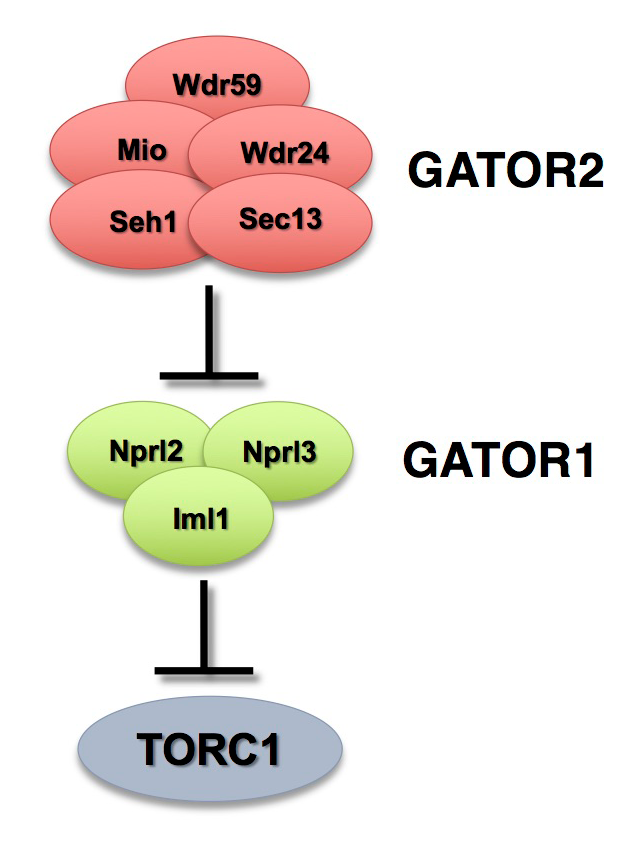
The Target of Rapamycin Complex 1 (TORC1) regulates cell growth and metabolism in response to many inputs, including amino acid availability and intracellular energy status. In the presence of sufficient nutrients and appropriate growth signals, the Ragulator and the Rag GTPases (a complex that regulates lysosomal signaling and trafficking) target TORC1 to lysosomal membranes, where TORC1 associates with its activator, the small GTPase Rheb. Once activated, TORC1 is competent to phosphorylate its downstream targets. The GATOR (Gap activity towards Rags) complex is an upstream regulator of TORC1 activity. The GATOR complex consists of two subcomplexes (Figure 1): the GATOR1 complex inhibits TORC1 activity in response to amino-acid starvation and is a trimeric protein complex consisting of the proteins Nprl2, Nprl3, and Iml1. Evidence from yeast and mammals indicates that the components of the GATOR1 complex function as GTPase–activating proteins (GAP), which inhibit TORC1 activity by inactivating the Rag GTPases. Notably, Nprl2 and Iml1 are tumor-suppressor genes, and mutations in Iml1, known as DEPDC5 in mammals, are a leading cause of hereditary epilepsy.
Our work, as well as that of others, found that the GATOR2 complex activates TORC1 by opposing the TORC1–inhibitory activity of GATOR1. Intriguingly, computational analysis indicates that the GATOR2 components Mio and Seh1, as well as several other members of the GATOR2 complex, have structural features consistent with coatomer proteins and membrane-tethering complexes. In line with the structural similarity to proteins that influence membrane curvature, we showed that three components of the GATOR2 complex, Mio, Seh1, and Wdr24, localize to the outer surface of lysosomes, the site of TORC1 regulation. However, how GATOR2 inhibits GATOR1 activity, thus permitting robust activation of TORC1, remains unknown. Additionally, the role of the GATOR1 and GATOR2 complexes in both the development and physiology of multicellular animals remains poorly defined. Over the past year, we used molecular, genetic, and cell-biological approaches to define the in vivo functions of the GATOR1 and GATOR2 complexes in both Drosophila and mammals. Additionally, we examined how the regulation of metabolism during oogenesis by the GATOR1 complex impacts oocyte development and the maintenance of oocyte quality.
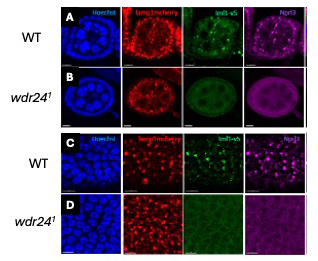
Figure 2. GATOR1 is absent from lysosomes in GATOR2 mutants.
In wild-type (A) nurse cells and (B) follicle cells of the ovary, the GATOR1 components Iml1 (purple) and Nprl3 (green) colocalize with the lysosomal marker Lamp1. In contrast, in wdr241 mutant (C) nurse cells and (D) follicle cells, GATOR1 components are not concentrated on lysosomes.
Figure 2. GATOR1 is absent from lysosomes in GATOR2 mutants.
In wild-type (A) nurse cells and (B) follicle cells of the ovary, the GATOR1 components Iml1 (purple) and Nprl3 (green) colocalize with the lysosomal marker Lamp1. In contrast, in wdr241 mutant (C) nurse cells and (D) follicle cells, GATOR1 components are not concentrated on lysosomes.
The GATOR2 complex maintains lysosomal-autophagic function by inhibiting the degradation of MiT/TFE transcription factors.
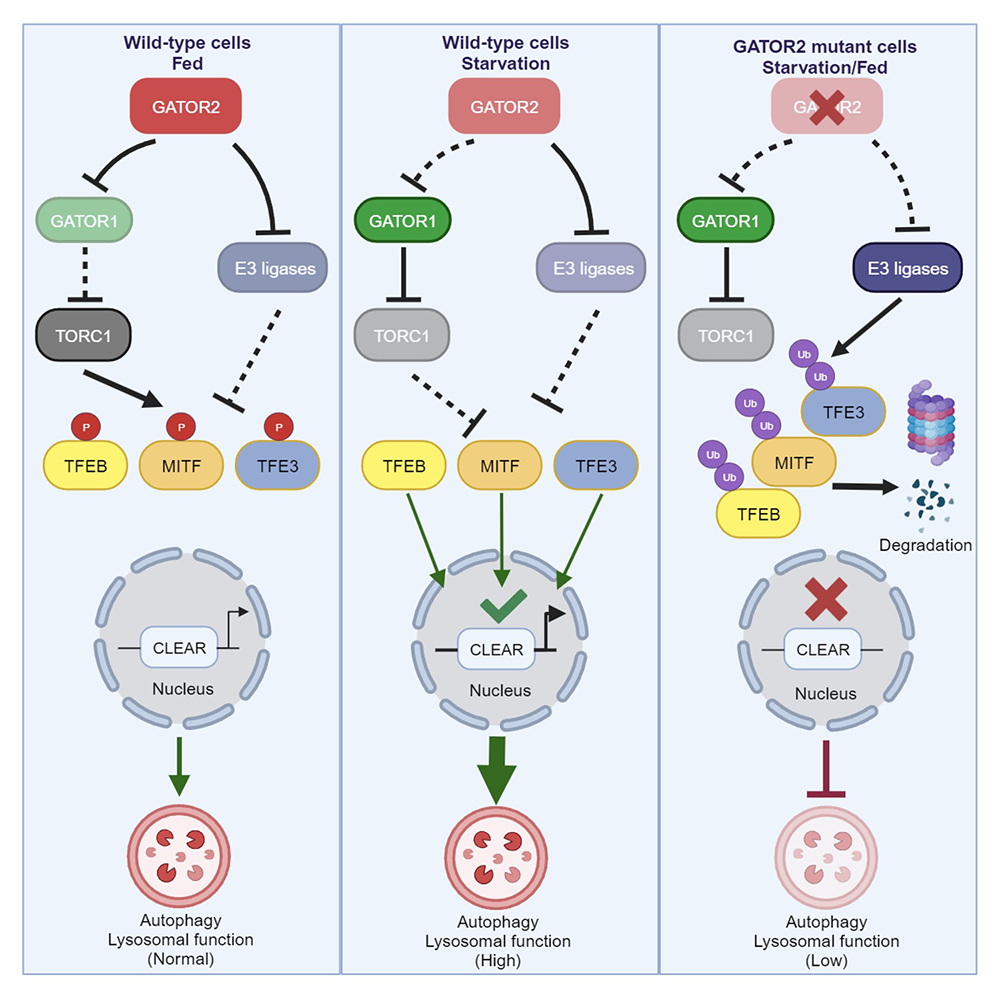
The lysosomal-autophagic system regulates the balance between anabolism and catabolism, which is essential to metabolic homeostasis. During periods of nutrient stress, cells upregulate lysosomal function and the autophagic process to promote the catabolic breakdown of macromolecules, such as proteins and lipids, to provide building blocks for cells to reuse. The MiT/TFE family of transcription factors promotes lysosomal-autophagic function by stimulating the transcription of numerous genes related to the lysosomal-autophagic system, including lysosomal-lumen enzymes, V-ATPase, and ATG proteins. A large body of work has uncovered a detailed mechanism for how TORC1 functions to inhibit the activation of MiT/TFE proteins through multiple mechanisms, including cytoplasmic retention and proteolytic degradation. These regulatory pathways allow MiT/TFEs to transcribe the genes required to promote catabolic metabolism during periods of nutrient stress, when TORC1 activity is low.
We discovered that the GATOR2 complex has independent roles in TORC1 regulation and MiT/TFE protein protection and is thus central to coordinating cellular metabolism with the control of the lysosomal-autophagic system. We found that, in cells with deletions of the GATOR2 subunits WDR24, MIOS, or SEH1, but not WDR59, members of the MiT/TFEs family of transcription factors TFEB, TFE3, and MITF are ubiquitylated and degraded by the proteasome. Low levels of MiT/TFEs result in a reduction in the transcription of a wide range of lysosomal-autophagic genes, including V-ATPase subunits, lysosomal enzymes (Cathepsins), structure proteins (LAMPs and LAMTORs), and several ATG genes (ATG9), causing a significantly lowered lysosomal pH and a systematic failure in digestion of lysosomal cargoes. We identified a trio of E3 ligases, HERC2, UBE3A, and STUB1, that target MiT/TFEs for ubiquitin-mediated degradation in GATOR2 knockout (KO) cells. We also found that, in autophagy-addicted pancreatic ductal adenocarcinoma (PDAC) cells, the GATOR2 complex is essential for maintaining PDAC cell proliferation, invasion, and malignancy by fulfilling its dual roles in cell metabolism, i.e., activation of TORC1 and protection of MiT/TFEs.
To complement the above studies, we recently knocked down the GATOR2 component WDR24 in two kidney-cancer cell models, UOK257 and UOK124. The UOK 257 cell line was derived from a human renal carcinoma of an individual with Birt-Hogg-Dubé (BHD) syndrome, which results from a mutation in the FLCN gene; FLCN regulates lysosome function by promoting the TORC1–dependent phosphorylation and cytoplasmic sequestration of TFEB. In the absence of FLCN, TFEB is not recruited to the lysosome but instead enters the nucleus and is active. The UOK124 cell line was derived from a Xp11.2 translocation renal cell carcinoma (tRCC). Xp11.2 tRCCs represent an aggressive type of kidney cancer resulting from genomic rearrangements of TFE3 that produce oncogenic TFE3 fusion proteins. Our preliminary data indicate that WDR24 knockdown results in markedly lowered levels of TFEB and TFE3 in UOK257 and UOK124 tRCC cells. We will continue to analyze how GATOR2 impacts these two cancer cell lines using strategies similar to those we employed to follow MiT/TFE levels and function in HeLa and PDAC cells. Our data indicate that the GATOR2 complex acts upstream of the TORC1 regulation of MiT/TFE activity. Thus, the GATOR2 complex may be an attractive target for chemotherapeutic intervention.
Publications
- The GATOR2 complex maintains lysosomal-autophagic function by inhibiting the protein degradation of MiT/TFEs. Mol Cell 2024 84:727-743
- Unveiling GATOR2 function: novel insights from Drosophila research. Cells 2024 13(21):1795
Collaborators
- Ryan Dale, MS, PhD, Bioinformatics and Scientific Programming Core, NICHD, Bethesda, MD
- Thomas Gonatopoulos-Pournatzis, PhD, Functional Transciptomics Section, NCI, Frederick, MD
- Marston Linehann, MD, Center for Cancer Research, NCI, Bethesda, MD
- Yingbiao Zhang, PhD, Institute for Translational Medicine, The Affiliated Hospital of Qingdao University, Qingdao, China
Contact
For more information, email mlilly@helix.nih.gov or visit https://www.nichd.nih.gov/research/atNICHD/Investigators/lilly.