Three-Dimensional Organization of the Genome as a Determinant of Cell-Fate Decisions

- Pedro Rocha,
PhD, Head, Unit on Genome Structure and Regulation - Nina Kopitchinski, BS, Lab Manager
- Manon Baudic, PhD, Postdoctoral Fellow
- Shreeta Chakraborty, PhD, Postdoctoral Fellow
- Joyce J. Thompson, PhD, Postdoctoral Fellow
- Zhenyu Zuo, PhD, Postdoctoral Fellow
- Samantha Bunner, BS, Postbaccalaureate Fellow
- Jack Waite, BA, Postbaccalaureate Fellow
Packing two meters of DNA inside ten-micron nuclei leads to a highly crowded environment and is thus an extraordinary feat of compaction and organization, because cellular processes such as DNA repair, replication, and transcription must occur within such a dense setting. We have all seen how electric cables become invariably entangled and non-functional inside our drawers. Cells face the same problems and have therefore developed mechanisms that organize the physical structure of the genome and ensure its regulation and integrity. Our lab studies how genome folding not only fits DNA within the constraints of the nuclear space but also is used as a critical mechanism of gene regulation. This is particularly important during development because dysregulation of gene expression causes severe birth defects.
We have shown that variations of just a few nucleotides of DNA sequence can disrupt genome architecture, dysregulate gene expression, and severely compromise mammalian development. In addition, some of the animal models we developed are being used to test therapies for defects caused by abnormal genome structure. While centered on embryogenesis, our work will shed light on how cell type–specific transcriptional programs are established, with broad implications for cell-fate decisions in disease contexts such as cancer.
Representative image of the lab's research
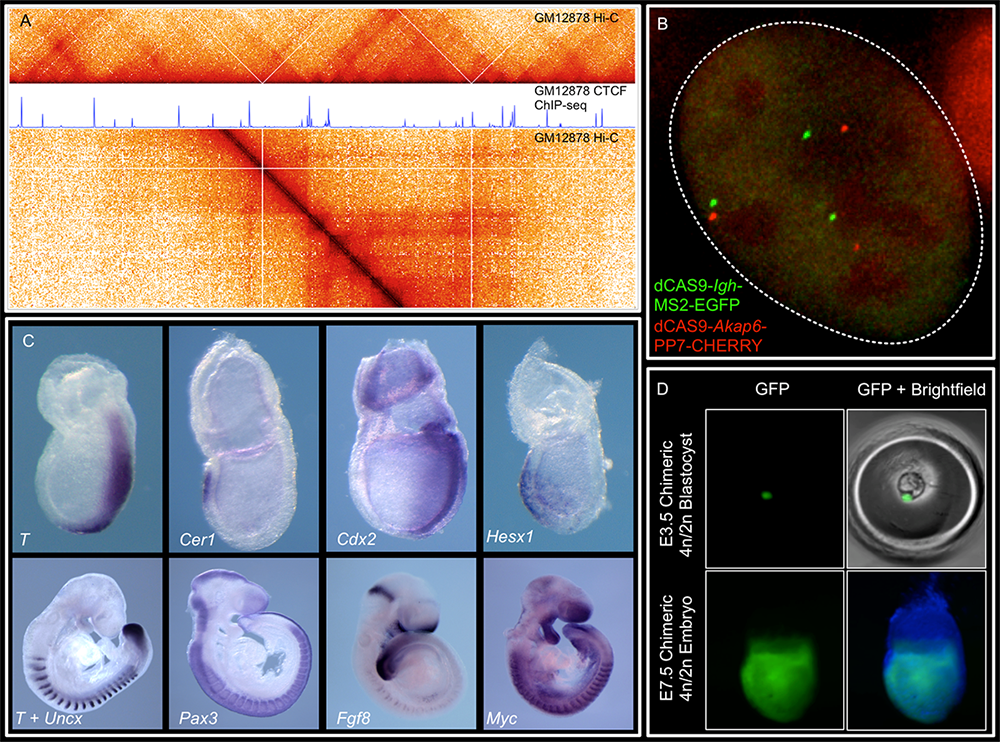
We combine imaging techniques in both fixed and living cells with sequencing-based genomic techniques that assess DNA–DNA interactions.
A. Hi-C (genomic analysis technique) and CTCF ChIP-Seq of GM1278 cells (a standard cell line), which allow characterization of chromatin structure and identification of binding sites of an important architectural protein
B. dCAS9 MCP-EGFP (MS2-MCP is an imaging system widely used to study mRNA spatial distribution in living cells) and PCP-CHERRY (coat protein with red fluorescent protein tag) live imaging of the Igh and Akap6 (A-kinase anchoring protein 6) loci: the mouse embryo is an unparalleled system in mammalian biology for understanding how tissue-specific gene expression is achieved.
C. Whole-mount in situ hybridization for patterning markers in mid and late gastrulating embryos
D. Tetraploid aggregation with GFP (green fluorescent protein)–labeled ES (embryonic stem) cells allows generation of fully ES cell–derived embryos.
Representative image of the lab's research
We combine imaging techniques in both fixed and living cells with sequencing-based genomic techniques that assess DNA–DNA interactions.
A. Hi-C (genomic analysis technique) and CTCF ChIP-Seq of GM1278 cells (a standard cell line), which allow characterization of chromatin structure and identification of binding sites of an important architectural protein
B. dCAS9 MCP-EGFP (MS2-MCP is an imaging system widely used to study mRNA spatial distribution in living cells) and PCP-CHERRY (coat protein with red fluorescent protein tag) live imaging of the Igh and Akap6 (A-kinase anchoring protein 6) loci: the mouse embryo is an unparalleled system in mammalian biology for understanding how tissue-specific gene expression is achieved.
C. Whole-mount in situ hybridization for patterning markers in mid and late gastrulating embryos
D. Tetraploid aggregation with GFP (green fluorescent protein)–labeled ES (embryonic stem) cells allows generation of fully ES cell–derived embryos.
Research goals
Studying gene regulation is critical to understanding how disruptions in developmental gene expression lead to birth defects and to enable earlier diagnosis and potential intervention strategies. We investigate how genome folding not only fits the long DNA fiber within the constraints of the nuclear space but also how it is used as a critical mechanism of gene regulation during cell-fate specification. This is a key fundamental question because transcription of many genes required for healthy development is controlled by non-coding regulatory elements, known as enhancers, that can be found at large genomic distances from their target genes. Our long-term scientific goal is to decipher how, within a crowded nucleus, distal enhancers are guided specifically to their target genes and which mechanisms prevent off-target activation.
1. Identification and characterization of DNA–regulatory elements that facilitate or restrict long-distance activation
Such elements may shape genome folding globally and indirectly modulate enhancer interactions, for example DNA motifs bound by CTCF (ubiquitously expressed zinc finger DNA–binding protein). Alternatively, some sequences may directly tether contacts with enhancers. CpG islands (unique DNA sequences that are characterized by their abundance of CpG dinucleotides and are typically found in the promoter regions of genes) have been proposed to do this in vertebrates. We use unbiased and candidate-based functional approaches to discover novel elements and characterize their function.
2. Discovery of regulatory complexes that shape enhancer-promoter contacts
Identifying motifs will be the first step to pinpoint the regulatory complexes that establish enhancer contacts and how they induce transcription. We will focus on transcription factors as well as large co-regulatory complexes, and the role of the polymerase machinery in setting interactions between regulatory elements.
3. Improvement of disease outcomes of developmental disorders
Our aims are to identify genomic regions as well as nuclear factors that may be hypersensitive to sequence variation. Not only will this enable early diagnosis, but may also help design targeted therapies. The development of animal models can then be used to directly test such therapies.
Approach
We generate allelic series of genetically engineered mouse models with different types of perturbations of chromatin structure, and employ advanced genomics techniques to molecularly phenotype these mutants. We complement in vivo work with cell culture–based differentiation, where we employ our modifications of massively parallel reporter assays to probe the functional role of different regulatory elements in controlling gene expression under distinct developmental and genomic contexts.
CTCF loops promote and repress enhancer contacts.
These loops form through the ability of the cohesin complex to extrude DNA and of CTCF (CCCTC–binding factor) to retain extruding cohesin. CTCF loops (key regulatory proteins of three-dimensional genome organization) can facilitate contacts between enhancers and genes but also ensure that enhancers cannot activate genes outside of their loop. However, it is not yet clear how these models are generalizable to all genes and how the impact of CTCF–cohesin loops on gene regulation may be affected by different types of genes, enhancers, and loops.
Transcriptional insulation by CTCF loops is modulated by enhancer type and boundary strength.
SOX2 is a transcription factor important for epiblast specification, neurogenesis, and tracheo-esophageal separation. Its locus is bound by CTCF directly upstream of its promoter, which suggest that CTCF–cohesin loops could be used to recruit its different tissue-specific enhancers. Surprisingly, we showed [Reference 2] that Sox2 recruits its enhancers through CTCF–independent mechanisms. We also generated an extensive series of mouse mutants with insertions of ectopic loops between Sox2 and its enhancers. By comparing different types of insertions, we determined that the highest transcriptional insulation is achieved when extruding cohesin complexes are blocked by CTCF from both directions of an active enhancer-promoter pair.
In contrast to the epiblast and to neural tissues, Sox2 was undetectable in the anterior foregut of mutants carrying the strongest boundaries, and these animals fully phenocopied loss of SOX2 in this tissue. The blastocyst and neural Sox2 enhancers are found within high-density clusters of transcription factor motifs, and they recruit high levels of coregulatory complexes. In contrast, in the anterior foregut, Sox2 enhancers are spread through large genomic stretches at lower density. We therefore proposed that these different enhancer configurations determine the ability of enhancer-promoter interactions to bypass CTCF loops.
Loss of a single CTCF motif can severely dysregulate gene expression and mouse development.
We hypothesized that in loops with multiple developmental regulators, CTCF could be particularly important to ensure tight tissue-specific expression. To test this, we studied a 63kb loop harboring three essential FGF (fibroblast growth factor) ligand genes: Fgf3, Fgf4, and Fgf15. Removing the most centromeric boundary (C1–C4) of the domain led to striking lethal phenotypes. In fact, this is the first demonstration that heterozygous loss of a CTCF boundary can cause fully penetrant perinatal lethality. These embryos developed encephalocele, together with ectopic FGF expression in the brain. Surprisingly, we showed that loss of the single CTCF motif (C1) oriented towards Ano1 (calcium-activated chloride channel) fully recapitulates these phenotypes but deleting the three motifs pointing to the FGF genes (C2–C4) does not. This study [Reference 1] revealed that sequence variants, as small as a single CTCF motif, at particular domain boundaries can have a surprisingly outsized effect and must be considered as potential sources of gene dysregulation in development and disease. This is the first mouse model of encephalocele, a birth defect that affects 1:10,000 human births. We are currently using it to investigate the etiology of this disease and potential therapies.
Rewiring of regulatory interactions
As cells progress through their differentiation trajectories to become more and more specified, regulatory interactions between enhancers and promoters have to be remodeled and re-established. We are very interested in understanding the dynamics of this reorganization, the mechanisms that drive it, and how its importance to specification of cell fates may change throughout development. For example, during specification of the primitive endoderm, we showed that the co-bound state of NANOG (a transcription factor that helps embryonic stem cells maintain pluripotency) and GATA6 (a zinc finger transcription factor that is important in the endodermal differentiation of organ tissues) in ICM (inner cell mass, part of the blastocyst) cells is followed by NANOG eviction and repression of epiblast genes [Reference 3]. This is accompanied by quick remodeling of chromatin and enhancer-promoter contacts. This dynamic rewiring of the genome is not unique to the blastocyst. We recently showed that a specialized cell type of our immune system, dendritic cells, also rearrange their genome throughout differentiation [Reference 4].
Additional Funding
- DIR Scientific Director's Award
Publications
- Deletion of a single CTCF motif at the boundary of a chromatin domain with three FGF genes disrupts gene expression and embryonic development. Dev Cell 2025 60:1838-1853
- Enhancer-promoter interactions can bypass CTCF-mediated boundaries and contribute to phenotypic robustness. Nat Genet 2023 55(2):280-290
- Extensive co-binding and rapid redistribution of NANOG and GATA6 during emergence of divergent lineages. Nat Commun 2022 13:4257
- Chromatin structure undergoes global and local reorganization during murine dendritic cell development and activation. Proc Natl Acad Sci USA 2022 119:e2207009119
- MLL3/MLL4 methyltransferase activities control early embryonic development and embryonic stem cell differentiation in a lineage-selective manner. Nat Genet 2023 55(4):693-705
Collaborators
- Bradley Bernstein, MD, PhD, Broad Institute and Dana-Farber Cancer Institute, Cambridge, MA
- Stefan Feske, MD, New York University School of Medicine, New York, NY
- Daniel Herranz, PharmD, PhD, Cancer Institute of New Jersey, Rutgers University, New Brunswick, NJ
- Claire Le Pichon, PhD, Unit on the Development of Neurodegeneration, NICHD, Bethesda, MD
- Mark Lewandoski, PhD, Cancer and Developmental Biology Laboratory, Center for Cancer Research, NCI, Frederick, MD
- Timothy Petros, PhD, Unit on Cellular and Molecular Neurodevelopment, NICHD, Bethesda, MD
- Achim Werner, PhD, Stem Cell Biochemistry Unit, NIDCR, Bethesda, MD
Contact
For more information, email pedro.rocha@nih.gov or visit https://www.nichd.nih.gov/research/atNICHD/Investigators/rocha.