Decoding Developmental Signaling in Vertebrate Embryos

- Katherine W. Rogers,
PhD, Head, Unit on Developmental Signaling - William K. Anderson, BS, Research Technician
- Caitlin Donahue, PhD, Postdoctoral Fellow
- Leanne E. Iannucci, PhD, Postdoctoral Fellow
- Selvaraj Velanganni, PhD, Postdoctoral Fellow
- Catherine E. Campbell, BS, NIH-Johns Hopkins University Graduate Partnership Program Student
- Matthew T. Monaghan, BS, Postbaccalaureate Fellow
- Micaela R. Murphy, BS, Postbaccalaureate Fellow
During embryonic patterning a single cell gives rise to a complex adult containing stereotypically arranged cells, tissues, and organs. Disruptions to this process can lead to congenital anomalies including structural birth defects. Decades of research have demonstrated that patterning is orchestrated by gradients of signaling molecules that control the behavior of embryonic cells. Signals regulate cell division, migration, and fate decisions, ultimately coordinating the development of healthy organisms. However, it has been challenging to dissect the specific mechanisms by which signaling pathways execute their many effects in developing embryos. The goal of the Unit on Developmental Signaling (UDS) is to understand how signaling orchestrates embryonic patterning.
The UDS uses a two-pronged approach to investigate signaling-mediated patterning. First, we create and share cutting-edge molecular optogenetic technology to experimentally manipulate signaling. Second, we apply these powerful approaches to investigate in vivo how signaling is decoded into the molecular responses that underpin embryogenesis. Molecular optogenetic strategies provide precise spatio-temporal control over biological processes, opening new research avenues. We use the zebrafish embryo for its many advantages: these externally fertilized, translucent embryos are ideal for light-based optogenetic and imaging experiments; a wide range of robust molecular, sequencing, and genetic methods are available; and decades of research have established zebrafish as a powerful model of vertebrate development. To decode signaling-based fate decisions, we optogenetically manipulate signaling in zebrafish embryos and use: 1) microscopy and optics to investigate how signaling gradients convey spatial patterning information in developing tissues; and 2) sequencing-based approaches (RNA/ATAC-Seq) to determine how signaling dynamics are interpreted at the levels of chromatin and gene expression.
Developing novel molecular optogenetic technology
We developed and characterized a suite of zebrafish-optimized optogenetic tools to reversibly activate three key developmental signaling pathways: FGF, BMP, and Nodal (Figure 1). This toolkit enables tunable, wavelength-dependent, spatially localized signaling activation in zebrafish embryos. We are also developing transgenic zebrafish lines expressing these signaling activators. We use this powerful optogenetic platform to investigate the developmental biology questions outlined in the following sections.
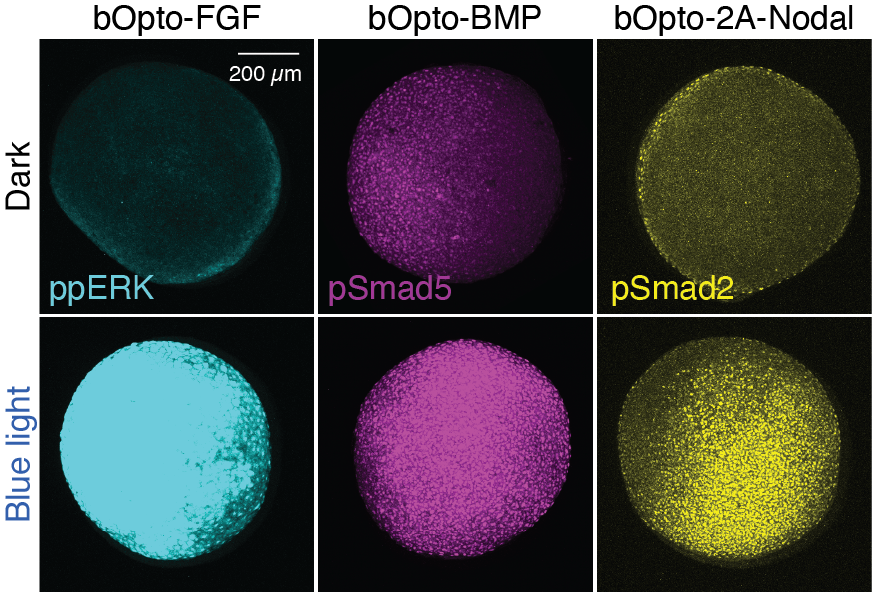
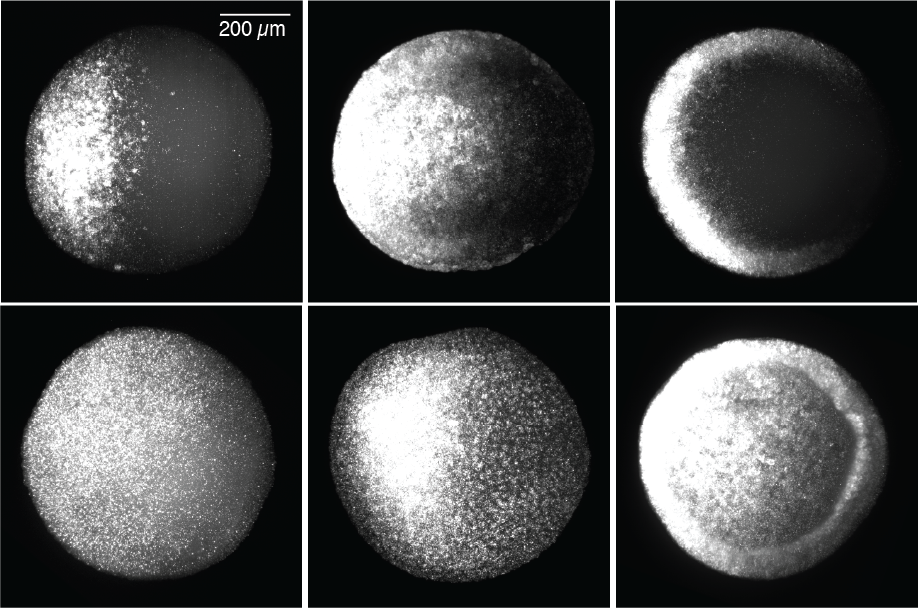
Figure 1. Zebrafish-optimized optogenetic signaling activator toolkit
Gastrulating zebrafish embryos expressing optogenetic activators of FGF, BMP, and Nodal signaling were exposed to blue light (455 nm) for 30 min (bottom row) or maintained in dark conditions (top row). Immunofluorescence staining for phosphorylated signaling effectors demonstrates blue light–mediated signaling activation in vivo.
Figure 1. Zebrafish-optimized optogenetic signaling activator toolkit
Gastrulating zebrafish embryos expressing optogenetic activators of FGF, BMP, and Nodal signaling were exposed to blue light (455 nm) for 30 min (bottom row) or maintained in dark conditions (top row). Immunofluorescence staining for phosphorylated signaling effectors demonstrates blue light–mediated signaling activation in vivo.
The advancement of molecular optogenetics has already yielded important insights in developmental biology and has promising applications in other fields, including regeneration and tissue engineering. However, there is currently a relative dearth of molecular optogenetic technology available in zebrafish, despite this translucent vertebrate model’s suitability for these approaches. The UDS is developing novel molecular optogenetic strategies to expand the repertoire of these valuable tools. We are working to build optogenetic technology to activate additional signaling pathways, to create reversible signaling inhibitors, and to develop tools responding to different wavelengths for orthogonal manipulation. To increase the reproducibility and ease of optogenetic experiments in zebrafish, we are continuing to generate transgenic lines expressing these tools.
What information is encoded in signaling gradients?
Developmental patterning relies on spatial signaling gradients. Classic models postulate that cells read out signaling-molecule concentration to make fate decisions, implying that specific spatial distributions are important for normal pattering. However, contemporary work suggests a more nuanced strategy, in which gradients may instead function by engaging a patterning network, requiring only spatial signaling asymmetry instead of a precise gradient shape. It is also possible that both mechanisms, or others, are used depending on the patterning context. To determine the “error tolerance” of signaling gradients during patterning, we will eliminate endogenous signaling and use our optogenetic toolkit to impose custom signaling distributions in zebrafish embryos (Figure 2). By assessing patterning consequences, such an approach will provide insights into the information content contained in developmental signaling gradients.
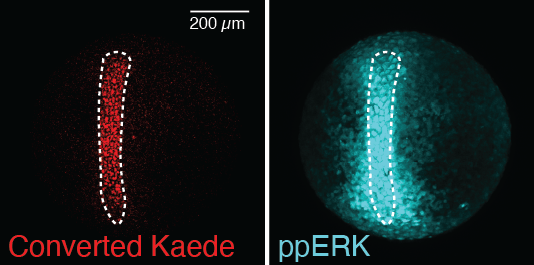
Figure 2. Localized signaling activation in vivo
A zebrafish embryo expressing the optogenetic FGF activator and the photo-convertible fluorescent protein Kaede was locally illuminated (white dotted line) with 405 and 445 nm confocal lasers. Immunofluorescence staining for the FGF–signaling effector ppERK shows localized signaling activation overlapping with photo-converted Kaede.
Figure 2. Localized signaling activation in vivo
A zebrafish embryo expressing the optogenetic FGF activator and the photo-convertible fluorescent protein Kaede was locally illuminated (white dotted line) with 405 and 445 nm confocal lasers. Immunofluorescence staining for the FGF–signaling effector ppERK shows localized signaling activation overlapping with photo-converted Kaede.
DNA–level mechanisms of dynamic signaling interpretation
Diverse spatio-temporal gene expression is necessary for healthy embryonic development (Figure 3). Gene expression is regulated by signaling, which is itself spatio-temporally dynamic. Each gene is thought to uniquely interpret signaling inputs including amplitude, duration, and combinations, leading to distinct expression profiles. This interpretation ultimately takes place at the level of DNA, through differential interactions with transcriptional regulators and chromatin changes. Yet the gene-specific input/output relationships between signaling, chromatin accessibility, and gene expression have not been systematically characterized during vertebrate development. Using our optogenetic toolkit, we are working to determine the mechanisms by which developmental genes decode signaling dynamics and combinations by optogenetically manipulating these inputs and profiling DNA– and RNA–level responses. Our in vivo signaling manipulations enable a direct link between signaling input and responses, and will help explain how diverse spatio-temporal gene expression profiles are generated in developing vertebrates.
Additional Funding
- NICHD Career Development Award (to Caitlin Donahue)
- NICHD Career Development Award (to Leanne Iannucci)
Publications
- An optogenetic toolkit for robust activation of FGF, BMP, & Nodal signaling in zebrafish. bioRxiv 2025 2025.04.17.649426; preprint
- Optogenetic signaling activation in zebrafish embryos. J Vis Exp 2023 10.3791/65733
- Optogenetic investigation of BMP target gene expression diversity. eLife 2020 e58641
- Optogenetic approaches to investigate spatiotemporal signaling during development. In: Gradients and Tissue Patterning. Small S, Briscoe J, eds, 2020 37–77
- Dynamics of BMP signaling and distribution during zebrafish dorsal-ventral patterning. eLife 2017 e25861
Contact
For more information, email katherine.rogers@nih.gov or visit https://www.nichd.nih.gov/research/atNICHD/Investigators/rogers.