Molecular Mechanisms of Synapse Assembly and Homeostasis

- Mihaela Serpe,
PhD, Head, Section on Cellular Communication - Peter Nguyen, Biological Laboratory Technician
- Tae Hee Han, PhD, Staff Scientist
- Rosario Vicidomini, PhD, Research Fellow
- Shefali Goyal, PhD, Visiting Fellow
- Wen Chieh Hsieh, PhD, Visiting Fellow
- Debabrata Sinha, PhD, Visiting Fellow
- Emmet Benaryeh, BS, Postbaccalaureate Fellow
The purpose of our research is to understand the mechanisms of synapse development and homeostasis. The chemical synapse is the fundamental nervous-system communication unit that connects neurons to one another and to non-neuronal cells, and it is designed to mediate rapid and efficient transmission of signals across the synaptic cleft. Crucial to this function is the ability of a synapse to change its properties, so that it can optimize its activity and adapt to the status of the cells engaged in communication and/or to the larger network comprising them. Consequently, synapse development is a highly orchestrated process coordinated by intercellular communication between the pre- and postsynaptic compartments and by neuronal activity itself. Our long-term goal is to elucidate the molecular mechanisms that regulate assembly of functional synapses during development and their fine-tuning during maturation, plasticity, and homeostasis. We use a comprehensive set of approaches that include genetics, biochemistry, molecular biology, cell biology, super-resolution imaging, and electrophysiology recordings in live animals and in reconstituted systems. In recent studies, we also utilized single-cell RNA sequencing (scRNA-Seq) methodologies to describe various populations of neurons.
Because of its many advantages, we use the Drosophila neuromuscular junction (NMJ) as a model for glutamatergic synapse development and function. The fact that individual NMJs can be reproducibly identified from animal to animal and are easily accessible for electrophysiological and optical analysis makes them uniquely suited for in vivo studies on synapse assembly, growth, and plasticity. In addition, the richness of genetic manipulations that can be performed in Drosophila permits independent control of individual synaptic components in distinct cellular compartments. Importantly, the fly NMJ relies entirely on kainate-type receptors, a family of ionotropic glutamate receptors (iGluRs) that impact synaptic transmission and neuronal excitability in the mammalian central nervous system but remain poorly understood. Unlike in vertebrates, where in vivo studies of kainate receptors have been technically highly challenging, disruption of kainate receptor biology at the Drosophila NMJ triggers a wide range of morphological and behavioral phenotypes, from a complete loss of postsynaptic receptors and synaptic structures, which causes embryonic paralysis and developmental lethality, to defects in the assembly and maintenance of PSDs (postsynaptic densities), which generally induce locomotor deficits, and to subtle changes in postsynaptic composition and impairments in synaptic plasticity and homeostasis. Within this wide range of biological outcomes, investigations of Drosophila NMJ iGluRs have the potential to reveal new functions for iGluRs and new modalities of regulation. The Drosophila NMJ can thus be used to analyze and model defects in the structural and physiological plasticity of glutamatergic synapses, which are associated with a variety of human pathologies, from learning and memory deficits to autism. Drosophila has long served as a source of insight into human genetics, development, and disease. The basic discoveries of our laboratory makes in the fly will serve our overarching goal of understanding how chemical synapses are assembled and sculpted during development and homeostasis.
The gating properties of Drosophila NMJ glutamate receptors and their dependence on Neto
Historically, a major obstacle to understanding ionotropic glutamate receptor (iGluR) function was the inability to reconstitute functional Drosophila NMJ receptors in heterologous systems and dissect their properties. Since iGluRs' recruitment and synaptic stabilization depend on their gating properties, such functional analyses are critical for comprehending the compound in vivo phenotypes of mutants with perturbed synaptic activities. We previously discovered that an obligatory auxiliary protein, Neto, is absolutely required for clustering of postsynaptic iGluRs and NMJ functionality. Neto belongs to a family of highly conserved auxiliary proteins that share an ancestral role in the formation and modulation of glutamatergic synapses. Vertebrate Neto1 and Neto2 were shown to modulate the properties of selective iGluRs, in particular kainate-type receptors. The fly NMJ relies exclusively on kainate-type receptors, making this genetically powerful system completely dependent on Neto activities.
Drosophila neto encodes two isoforms, Neto-α and Neto-β, which share the extracellular and transmembrane domains but have distinct intracellular parts. Our discovery of Drosophila Neto was a key step towards the functional reconstitution of recombinant NMJ receptors. In previous work we found that, when expressed in Xenopus oocytes, NMJ iGluRs in complex with Neto form rapidly desensitizing Ca2+–permeable channels with low affinity for glutamate. In the absence of Neto, the NMJ iGluRs reach the cell surfaces but elicit no glutamate-gated currents. More recently, we expressed functional NMJ iGluR/Neto complexes in HEK293T cells (an immortalized cell line derived from human embryonic kidney cells), in which the biophysical properties of these receptors can be further dissected using outside-out patches and fast agonist perfusion. As in Xenopus oocytes, Neto is critical for the functional reconstitution of both types of postsynaptic glutamate receptor complexes (type-A: GluRIIA/C/D/E and type-B: GluRIIB/C/D/E): 100% of the patches (n over 100) yielded no currents in the absence of Neto proteins.
We compared the properties of recombinant receptors (expressed in HEK293T cells) with native extra-junctional and synaptic receptors of genetically controlled composition. We found that the type-A and type-B receptors have comparable deactivation rates, but type-B desensitizes four times faster than type-A. Neto isoforms differentially modulate these channels, adding further diversity to their functional repertoire. For example, Neto-α speeds up the desensitization of both channels, whereas Neto-β and Neto-ΔCTD (Neto lacking its C-terminal domain) confer similar kinetics. This confirms our earlier findings that Neto-β functions primarily as an organizing scaffold at the NMJ. Surprisingly, we found that deactivation is extremely fast and that the decay of synaptic currents resembles the rate of iGluR desensitization. This indicates that a) the clearance of glutamate from the synaptic cleft is slow, b) additional auxiliary or modulatory proteins are present and impact the gating of synaptic receptors, and c) desensitization plays a major role in determining the kinetics of synaptic transmission at the larval NMJ.
Single-channel kinetics and sub-conductance states
Our studies revealed that Drosophila Neto is not only required for postsynaptic glutamate receptor channel function but also increases the repertoire of channel properties. Using outside-out patch recordings and fast agonist application, we also captured single-channel events and revealed major differences between the postsynaptic type-A and type-B glutamate receptors and between Neto splice variants.
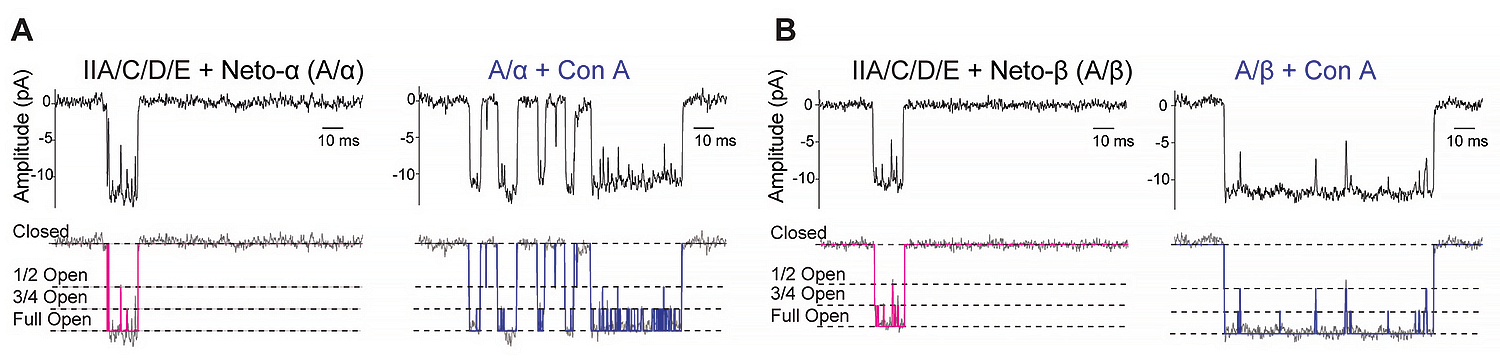
For example, inspection of single-channel records for type-A/Neto-α (A/α) and type-A/Neto-β (A/β) complexes revealed bursts of openings, during which the current fluctuated between the main state and well defined sub-conductance states with amplitudes of 75% and 50% (Figure 1); we did not observe openings to 25%, but it is possible that these might occur with lower concentrations of glutamate. The lectin concanavalin A (Con A) attenuates iGluR desensitization by binding at the interface between the amino-terminal domain and ligand-binding domain of iGluRs. Pretreatment with Con A dramatically increased single-channel activity without any change in single-channel conductance. We are currently expanding these single-channel kinetic analyses to gain mechanistic insights into the Neto-mediated modulation of postsynaptic glutamate receptors.
Figure 1. Analysis of single-channel kinetics for postsynaptic type-A glutamate receptors
A and B: filtered and idealized traces illustrating responses to 10 mM glutamate applied for 100 ms to outside-out patches from HEK293T cells transfected with A/α and A/β receptor complexes. Left: control patches. Right: patches obtained after treatment of HEK cells with 0.6 mg/mL Con A for 10 min; idealized traces, shown in magenta and blue, respectively, reveal the full open state and two subconductance states, half open and three-quarter open, as indicated by dotted lines.
Figure 1. Analysis of single-channel kinetics for postsynaptic type-A glutamate receptors
A and B: filtered and idealized traces illustrating responses to 10 mM glutamate applied for 100 ms to outside-out patches from HEK293T cells transfected with A/α and A/β receptor complexes. Left: control patches. Right: patches obtained after treatment of HEK cells with 0.6 mg/mL Con A for 10 min; idealized traces, shown in magenta and blue, respectively, reveal the full open state and two subconductance states, half open and three-quarter open, as indicated by dotted lines.
Previous work from our lab and others showed that Drosophila type-A and type-B receptors are recruited at postsynaptic densities (PSDs) by different mechanisms. Native type-B receptors desensitize faster that type-A; our recent kinetics studies captured different gating properties for recombinant and native type-A and type-B receptors. Also, previous work from our lab and others showed that the synaptic recruitment of type-A receptors depends on their gating properties and synaptic activity. In on-going single-channel kinetic experiments, we seek to determine the molecular basis for these differences.
Multilayer control of KaiR1D autoreceptor function by the auxiliary protein Neto
It has been long recognized that KARs (kainate receptors), operating as autoreceptors, provide a critical feedback mechanism that modulates neurotransmitter release and ensures stable neuronal network activities. Perturbations to this feedback have been associated with a wide range of neurological, psychiatric, and neurodegenerative disorders. At the Drosophila NMJ, KaiR1D (kainate-type ionotropic glutamate receptor subunit 1D) autoreceptor and Neto have been both positively implicated in the control of glutamate release. In the absence of KaiR1D or Neto-α, the basal neurotransmission levels are reduced to half of control levels. If Neto-α functions as a true auxiliary subunit for KaiR1D receptors, then its presence should be required for normal KaiR1D–mediated activities in the larval motor neurons. Indeed, we found that KaiR1D, even when in excess, cannot sustain normal neurotransmitter release in the absence of Neto-α. Neto-α may be required to modulate the gating properties of KaiR1D receptors, their presynaptic localization, or both.
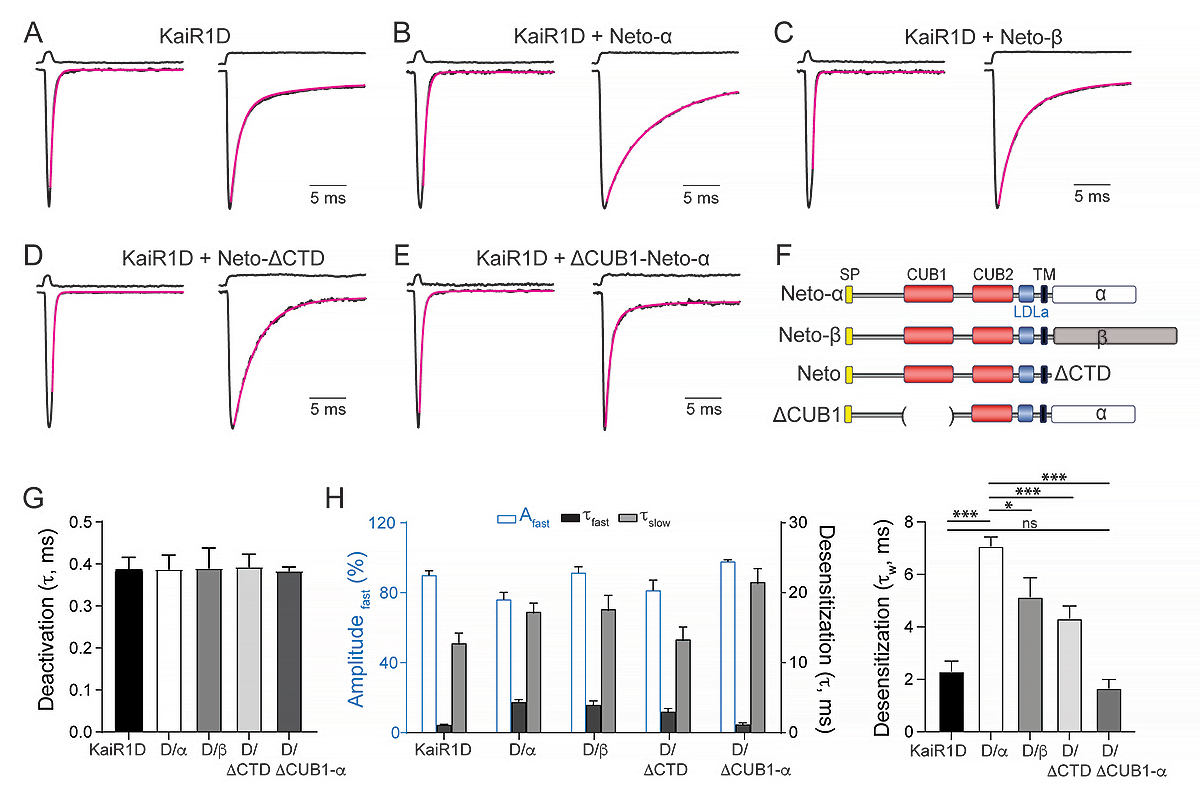
We first used outside-out patch-clamp recordings and fast ligand application to examine the biophysical properties of KaiR1D receptors expressed in HEK293T cells. KaiR1D forms functional homomeric ion channels with fast deactivation and desensitization time constants (Figure 2). Co-expression of Neto-α does not impact KaiR1D deactivation kinetics but slows the desensitization of KaiR1D channels three-fold. Neto-β has a smaller, two-fold impact. However, Neto-β cannot traffic to presynaptic locations and remains confined to the somato-dendritic compartment; its effect on KaiR1D desensitization is therefore irrelevant for presynaptic KaiR1D at the Drosophila NMJ; instead, Neto-β may modulate KaiR1D postsynaptic complexes elsewhere in the fly nervous system, for example in the adult eye. Neto-ΔCTD (Neto with no intracellular domain [i.e., a “minimal Neto”] but retains the highly conserved extracellular and transmembrane domains) also has a roughly two-fold impact on the desensitization of KaiR1D. By contrast, deletion of the CUB1 domain (CUB is an acronym for "complement C1r/C1s, Uegf, Bmp1," which mediates protein-protein interactions in various extracellular proteins) of Neto-α induced even faster desensitization of KaiR1D/ΔCUB1-Neto-α channels. This result is consistent with previous reports that the CUB1 domain of vertebrate Neto proteins stabilizes the structure and function of recombinant kainate receptor complexes in a heterologous expression system.
Figure 2. The effect of Neto variants on KaiR1D deactivation and desensitization
A–E: Responses to 10 mM glutamate applied for 1 ms (left traces) and 100 ms (right traces) to outside-out patches from cells transfected with KaiR1D alone (A) or together with Neto variants, Neto-α (B), Neto-β (C), Neto-ΔCTD (D) or ΔCUB1-Neto-α (E). Black lines show the average of 15–20 responses from one patch normalized and aligned to the peak. Red lines show decay of the responses fitted with the with the sum of one (deactivation) or two (desensitization) exponential functions; open tip junction currents measured at the end of the experiments are shown at the top of each panel. The holding potential was –60mV for all recordings.
F: Diagram of the Neto variants utilized.
G–H: Summary graph for deactivation (G) and desensitization time constants (H) for various KaiR1D/Neto complexes; the Afast (%) component for desensitization is shown in blue.
Statistical analyses are done with one-way ANOVA with post hoc Tukey's multiple comparisons. Data are represented as mean ± SEM. ***p < 0.001; *p < 0.05; ns, p > 0.05.
Figure 2. The effect of Neto variants on KaiR1D deactivation and desensitization
A–E: Responses to 10 mM glutamate applied for 1 ms (left traces) and 100 ms (right traces) to outside-out patches from cells transfected with KaiR1D alone (A) or together with Neto variants, Neto-α (B), Neto-β (C), Neto-ΔCTD (D) or ΔCUB1-Neto-α (E). Black lines show the average of 15–20 responses from one patch normalized and aligned to the peak. Red lines show decay of the responses fitted with the with the sum of one (deactivation) or two (desensitization) exponential functions; open tip junction currents measured at the end of the experiments are shown at the top of each panel. The holding potential was –60mV for all recordings.
F: Diagram of the Neto variants utilized.
G–H: Summary graph for deactivation (G) and desensitization time constants (H) for various KaiR1D/Neto complexes; the Afast (%) component for desensitization is shown in blue.
Statistical analyses are done with one-way ANOVA with post hoc Tukey's multiple comparisons. Data are represented as mean ± SEM. ***p < 0.001; *p < 0.05; ns, p > 0.05.
Neto-α and additional unknown modulators might also contribute to KaiR1D entering the axonal compartment and trafficking to presynaptic terminals in larval motor neurons. To eliminate the contribution of such unknown modulators and to test for a direct role for Neto in KaiR1D axonal distribution, in collaboration with Zheng Li, we set up an ex vivo experiment, transfecting recombinant Drosophila KaiR1D and Neto-α in 15-day in-vitro (DIV15) cultured primary rat hippocampal neurons and examining their subcellular distribution four days later (DIV19); the bright fluorescent reporter protein Venus was also introduced to visualize the transfected neurons.
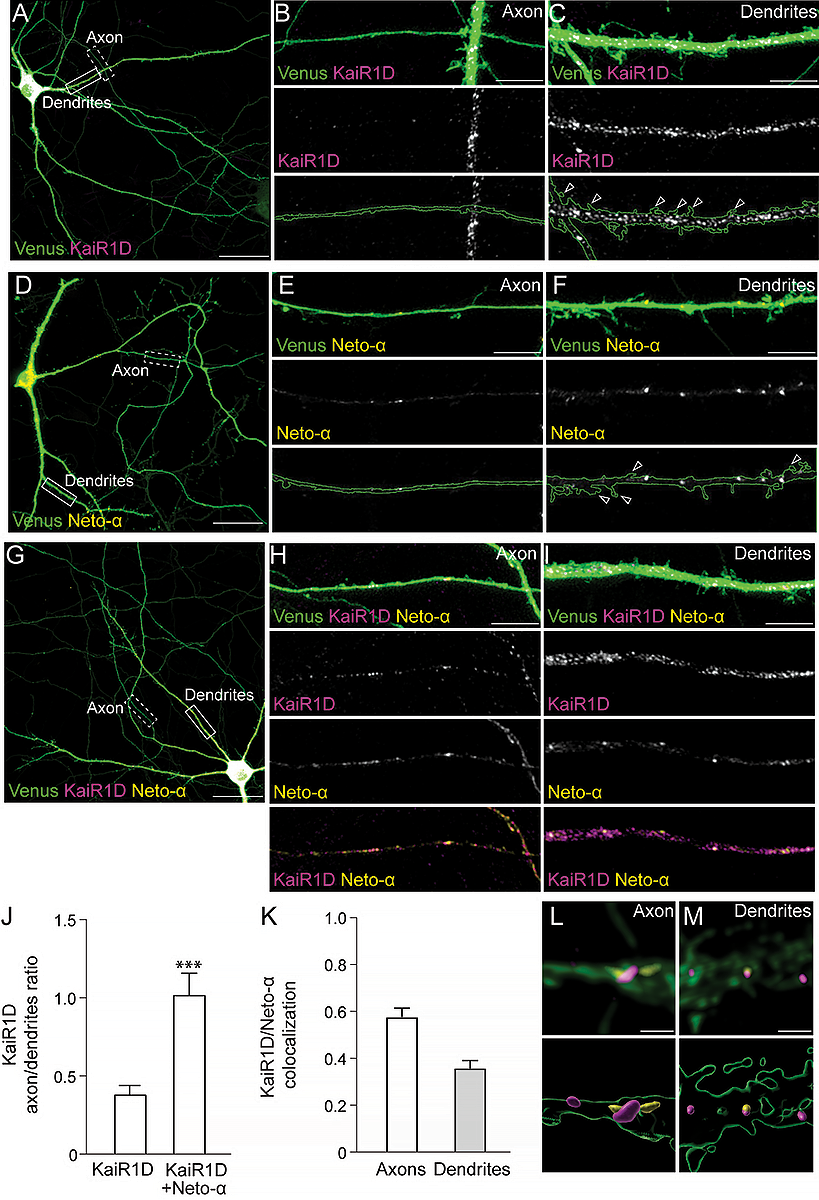
We found that, when expressed by itself, KaiR1D-Flag was marginally detectable in the axon; albeit, it accumulated as clear puncta in the somato-dendritic compartment (Figure 3). In contrast, Neto-α–positive puncta of various sizes were visible in both dendritic and axonal compartments. Co-expression of KaiR1D and Neto-α together did not change the distribution of Neto-α to neurites but had a significant effect on the presence of KaiR1D in the axonal compartment. This suggests that Neto-α promotes KaiR1D axonal localization. To confirm this apparent increase of axonal KaiR1D distribution and to account for variable transfection efficiency among different neurons/experiments, we quantified and normalized the density of KaiR1D puncta in axons versus dendrites in the absence or presence of Neto-α. Neto-α increased KaiR1D puncta density observed along the axons three-fold.
Figure 3. Neto-α promotes KaiR1D axonal distribution.
A–I: Confocal images of primary rat hippocampal neurons transfected (DIV15) and stained (DIV19) for KaiR1D (magenta), with Venus (green), and for Neto-α (yellow). Low magnification views (A, D, and I) capture the cell body as well as part of the axon and dendrites, shown at higher magnification in details (B–C, E–F, and H–I).
J: Quantification of KaiR1D puncta density in axon vs. secondary dendrites.
K: Colocalization between KaiR1D and Neto-α in axons and secondary dendrites.
L–M: 3D reconstitution of subcellular localization of KaiR1D and Neto-α along the axon (L) or in the dendrites (M).
Data are represented as mean ± SEM. Indicated P values are from unpaired two-tailed Mann-Whitney tests. ***p < 0.001.
Scale bars: 40 µm (A, D, and G), 5 µm (B, C, E, F, H, and I), and 1 µm (H' and I').
Figure 3. Neto-α promotes KaiR1D axonal distribution.
A–I: Confocal images of primary rat hippocampal neurons transfected (DIV15) and stained (DIV19) for KaiR1D (magenta), with Venus (green), and for Neto-α (yellow). Low magnification views (A, D, and I) capture the cell body as well as part of the axon and dendrites, shown at higher magnification in details (B–C, E–F, and H–I).
J: Quantification of KaiR1D puncta density in axon vs. secondary dendrites.
K: Colocalization between KaiR1D and Neto-α in axons and secondary dendrites.
L–M: 3D reconstitution of subcellular localization of KaiR1D and Neto-α along the axon (L) or in the dendrites (M).
Data are represented as mean ± SEM. Indicated P values are from unpaired two-tailed Mann-Whitney tests. ***p < 0.001.
Scale bars: 40 µm (A, D, and G), 5 µm (B, C, E, F, H, and I), and 1 µm (H' and I').
Interestingly, KaiR1D axonal delivery was not diminished in the presence of ΔCUB1-Neto-α and instead appeared enhanced compared with KaiR1D co-expressed with Neto-α. This indicates that the CUB1 domain is dispensable for KaiR1D axonal delivery. In contrast, a C-terminal–truncated Neto, Neto-ΔCTD, which lacks any intracellular domain, distributed to both axons and dendrites when transfected alone, but could no longer promote KaiR1D axonal delivery in co-expression experiments. These results indicate that the intracellular domain of Neto-α is critical for the distribution of KaiR1D to axons.
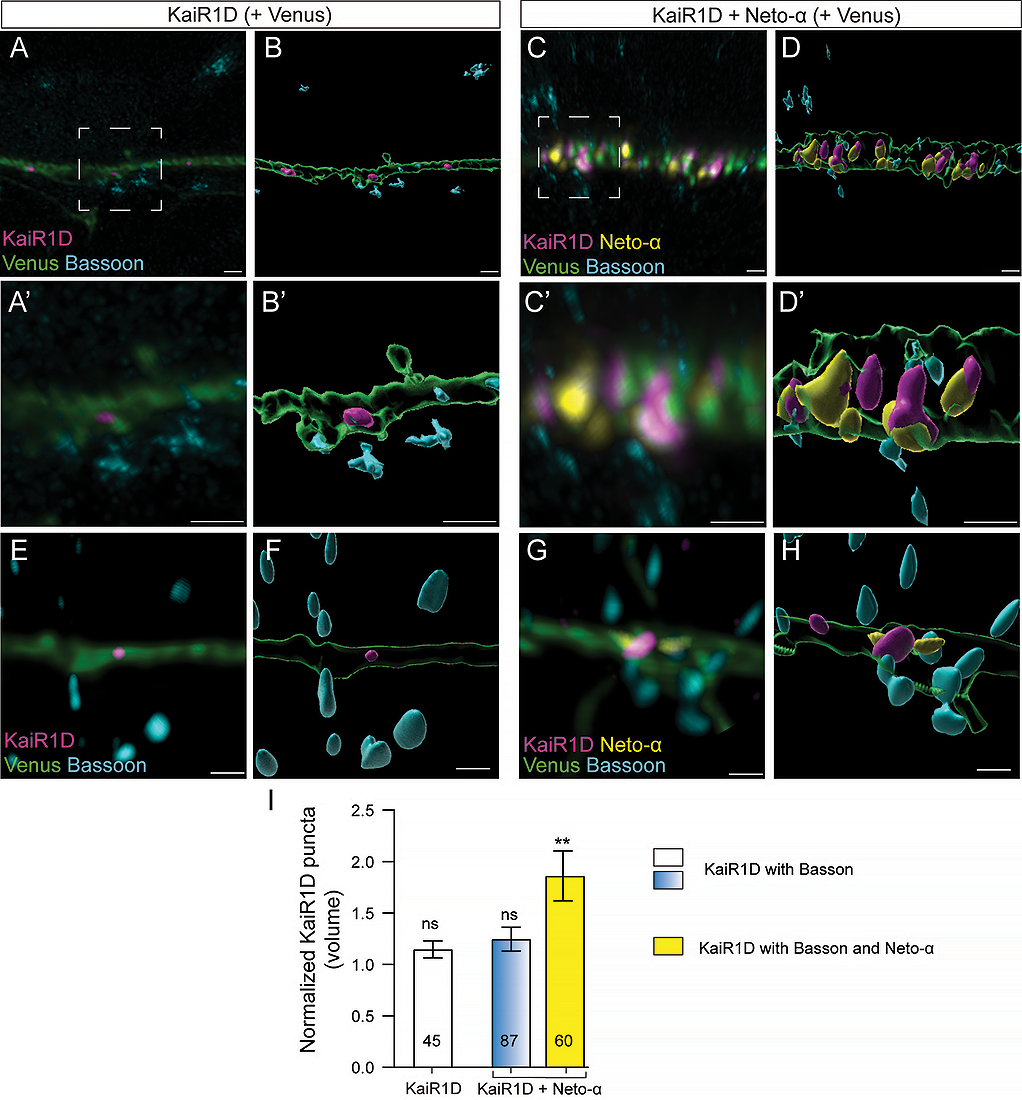
There was limited colocalization between KaiR1D and Neto-α in the dendrites. This may reflect the nature of these proteins, given that both Drosophila KaiR1D and Neto-α likely lack the protein features/interaction motifs required for recruitment and stabilization at vertebrate excitatory synapses. By contrast, a large proportion of axonal KaiR1D puncta colocalized with Neto-α signals (584 out of 889, n=32 cells). These results suggest that KaiR1D and Neto-α form complexes that traffic together and/or interact at specific locations along the axons. In flies as well as in vertebrate systems, Neto proteins and KARs stabilize each other at synaptic sites. Thus, it is possible that Drosophila KaiR1D and Neto-α colocalize at presynaptic sites when co-expressed in primary hippocampal neurons. In mammals, presynaptic locations are marked by Bassoon, a scaffolding protein involved in the active zone organization. Indeed, without Neto-α, we rarely observed accumulation of KaiR1D–positive puncta in the vicinity of Bassoon immunoreactivities (45 out of 703 puncta analyzed) (Figure 4). When co-transfected with Neto-α, the number of KaiR1D–positive puncta colocalized with Bassoon slightly increased (87 out of 1170), but, more importantly, puncta positive for both KaiR1D and Neto-α were significantly enlarged. These observations are consistent with stabilization of KaiR1D/Neto-α complexes at presynaptic sites. Together, our results indicate that Neto-α promotes KaiR1D axonal distribution and colocalizes with KaiR1D in the axonal compartment; nonetheless, KaiR1D and Neto-α may travel largely independently of each other to the dendrites.
Figure 4. KaiR1D and Neto-α accumulate at presynaptic sites.
Confocal images and 3D reconstitution of presynaptic contacts from primary rat hippocampal neurons transfected (DIV15) with KaiR1D alone (upper) or with Neto-α (lower) and stained (DIV19) with Venus (green), for Bassoon (cyan), for KaiR1D (magenta), and for Neto-α (yellow). When transfected without Neto-α, KaiR1D rarely enters the axonal compartment and does not colocalize with Bassoon. When transfected together, KaiR1D and Neto-α form complexes adjacent to Bassoon puncta. Data are represented as mean ± SEM. Indicated P values are from one-way ANOVA with Karuskal-Wallis multiple comparison test. **p < 0.01. Scale bar: 1 µm.
Figure 4. KaiR1D and Neto-α accumulate at presynaptic sites.
Confocal images and 3D reconstitution of presynaptic contacts from primary rat hippocampal neurons transfected (DIV15) with KaiR1D alone (upper) or with Neto-α (lower) and stained (DIV19) with Venus (green), for Bassoon (cyan), for KaiR1D (magenta), and for Neto-α (yellow). When transfected without Neto-α, KaiR1D rarely enters the axonal compartment and does not colocalize with Bassoon. When transfected together, KaiR1D and Neto-α form complexes adjacent to Bassoon puncta. Data are represented as mean ± SEM. Indicated P values are from one-way ANOVA with Karuskal-Wallis multiple comparison test. **p < 0.01. Scale bar: 1 µm.
Neto-α–mediated modulation of KaiR1D autoreceptor function is essential for normal basal neurotransmission.
Our in vitro and ex vivo studies indicate that Neto-α modulates both the function and axonal distribution of KaiR1D–containing autoreceptors. To address the question as to whether any of these Neto-α functions are essential for normal KaiR1D–dependent neurotransmitter release in vivo, we examined whether the Neto-α CUB1 and CTD domains are required for larval NMJ function, using in vivo rescue experiments.
Evoked junction potentials (EJPs) recorded at larval NMJ were reduced by half in absence of Neto-α (Figure 5). Neuronal overexpression of full-length Neto-α or Neto-ΔCTD, but not ΔCUB1-Neto-α, rescued the EJPs at neto-αnull NMJs. This difference could not be attributed to different protein levels or impaired traffic to presynaptic terminals, given that all Neto variants concentrated robustly at synaptic boutons. Instead, the inability of ΔCUB1-Neto-α to rescue basal neurotransmission in neto-αnull mutants indicates that this Neto variant cannot sustain KaiR1D function in vivo. This may reflect the extremely fast desensitization kinetics of KaiR1D/ΔCUB1-Neto-α channels and/or the inability of ΔCUB1-Neto-α to properly engage KaiR1D receptor complexes. If ΔCUB1-Neto-α engages KaiR1D inefficiently, then overexpression of ΔCUB1-Neto-α in motor neurons should have a minimal impact on neurotransmission; on the other hand, if ΔCUB1-Neto-α binds to KaiR1D and does not slow desensitization of these channels in vivo, then excess ΔCUB1-Neto-α should compete with endogenous Neto-α for KaiR1D and reduce the EJPs. We found that neuronal overexpression of ΔCUB1-Neto-α in an otherwise wild-type background reduces the basal transmission levels, suggesting that excess ΔCUB1-Neto-α displaces endogenous Neto-α from presynaptic KaiR1D/Neto-α complexes; the accumulation of fast desensitizing KaiR1D/ΔCUB1-Neto-α complexes then reduces neurotransmitter release. Together, these data demonstrate that Neto-α–dependent KaiR1D desensitization is a key element for maintaining normal basal neurotransmission at this synapse.
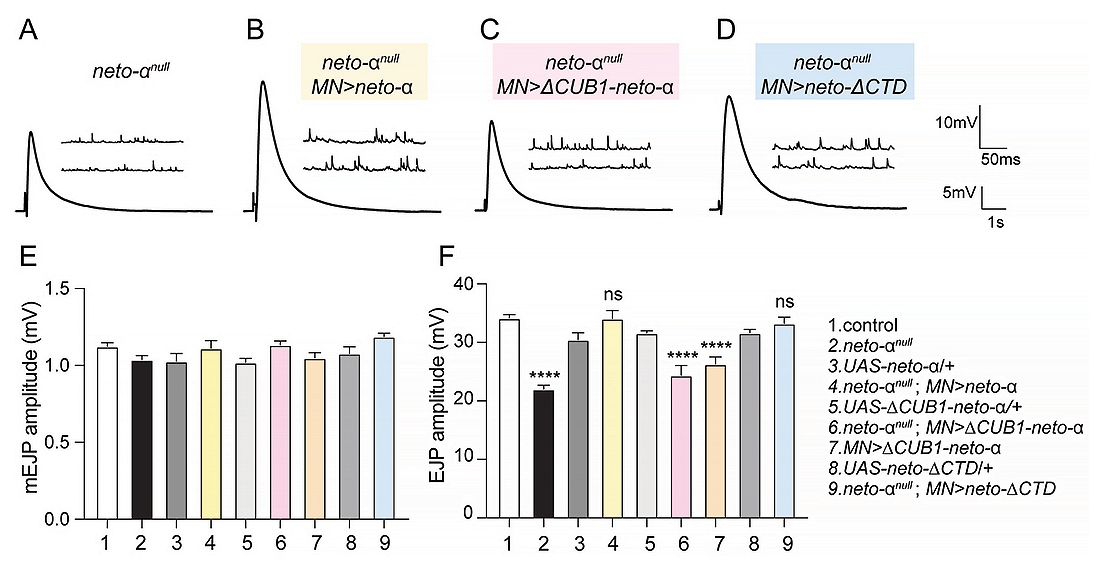
Figure 5. Neto-α CUB1 domain is essential for normal neurotransmission at larval NMJs (neuromuscular junctions).
A–D: Representative traces for mEJP (evoked junction potentials) and EJP recordings from muscle 6 (abdominal segments 3–4) of the indicated genotypes.
E–F: Quantification of mEJP and EJP amplitudes. Overexpression of Neto-α variants that slow KaiR1D desensitization rescues basal neurotransmission levels at neto-𝛼null NMJs, whereas Neto-α-ΔCUB1 does not, even though it promotes KaiR1D axonal delivery.
Statistical analyses are done with one-way ANOVA with post hoc Tukey's multiple comparisons. Data are represented as mean ± SEM.****p < 0.0001; ns, p > 0.05
Figure 5. Neto-α CUB1 domain is essential for normal neurotransmission at larval NMJs (neuromuscular junctions).
A–D: Representative traces for mEJP (evoked junction potentials) and EJP recordings from muscle 6 (abdominal segments 3–4) of the indicated genotypes.
E–F: Quantification of mEJP and EJP amplitudes. Overexpression of Neto-α variants that slow KaiR1D desensitization rescues basal neurotransmission levels at neto-𝛼null NMJs, whereas Neto-α-ΔCUB1 does not, even though it promotes KaiR1D axonal delivery.
Statistical analyses are done with one-way ANOVA with post hoc Tukey's multiple comparisons. Data are represented as mean ± SEM.****p < 0.0001; ns, p > 0.05
Our work has uncovered multiple roles for Neto-α in shaping the excitatory transmission; Neto-α secures the deployment of KaiR1D receptor complexes to axonal release sites and strengthens the signaling of individual channels. Such multi-layered control, choreographed through the functions of a single auxiliary subunit, ensures that KaiR1D activities are both potent and precisely localized. This Neto–integrated control mechanism parallels the multifunctionality known for AMPAR (ionotropic glutamate receptor) auxiliary proteins and indicates that Neto imparts an equally sophisticated regulatory complexity to kainate receptors. In future, it will be important to examine a role for Neto as a signaling hub that receives information about the network status and adjusts kainate receptor synaptic levels and channel properties, fine-tuning the amplitude of the synaptic output. In evolutionary terms, the presence of this multi-layered regulation of a kainate autoreceptor in Drosophila implies that coupling receptor function and localization may represent an ancient and conserved strategy to guarantee reliable neurotransmission.
Publications
- The gating properties of Drosophila NMJ glutamate receptors and their dependence on Neto. J Physiol 2024 602(24):7043-7064
- scRNA-seq data from the larval Drosophila ventral cord provides a resource for studying motor systems function and development. Dev Cell 2024 59(9):1210-1230
- Versatile nanobody-based approach to image, track and reconstitute functional Neurexin-1 in vivo. Nat Commun 2024 15:6068
Collaborators
- James Iben, PhD, Molecular Genetics Core, NICHD, Bethesda, MD
- Zheng Li, PhD, Section on Synapse Development Plasticity, NIMH, Bethesda, MD
- Dragan Maric, PhD, Flow and Imaging Cytometry Core, NINDS, Bethesda, MD
- Mark L Mayer, PhD, Section on Neurophysiology and Biophysics, NINDS, Bethesda, MD
- Felipe Opazo, PhD, Center for Biostructural Imaging of Neurodegeneration, Göttingen, Germany
Contact
For more information, email mihaela.serpe@nih.gov or visit https://www.nichd.nih.gov/research/atNICHD/Investigators/Serpe.