Regulatory Small RNAs and Small Proteins

- Gisela Storz,
PhD, Head, Section on Environmental Gene Regulation - Aixia Zhang, PhD, Staff Scientist
- Maciej M. Basczok, PhD, Postdoctoral Fellow
- Rajat Dhyani, PhD, Postdoctoral Fellow
- Chelsey R. Fontenot, PhD, Postdoctoral Fellow
- José Hernández-Valle, PhD, Postdoctoral Fellow
- Madison Jermain, PhD, Postdoctoral Fellow
- Janka J. Schmidt, PhD, Postdoctoral Fellow
- Shuwen Shan, PhD, Postdoctoral Fellow
- Narumon Thongdee, PhD, Postdoctoral Fellow
- Rilee D. Zeinert, PhD, Postdoctoral Fellow
- Aoshu Zhong, PhD, Postdoctoral Fellow
- Dennis X. Zhu, PhD, Postdoctoral Fellow
- Zachary Rich, BA, Graduate Student
- Amanda Brewer, BS, Postbaccalaureate Fellow
- Anna J. Bryant, BS, Postbaccalaureate Fellow
The group currently has two main interests: identification and characterization of small noncoding RNAs (sRNAs), and identification and characterization of small proteins of less than 50 amino acids. Both small RNAs and small proteins, also known as microproteins, have been overlooked because they are not detected in biochemical assays, and the corresponding genes are missed by genome annotation and are poor targets for genetic approaches. However, both classes of small molecules are being found to have important regulatory roles in organisms ranging from bacteria to humans.
Identification and characterization of small regulatory RNAs
During the past 25 years, we carried out several different systematic screens for small regulatory RNAs (sRNAs) in Escherichia coli. The screens included computational searches for conservation of intergenic regions and direct detection after size selection or co-immunoprecipitation with RNA–binding proteins. Most recently, we have been using deep sequencing approaches to map the 5′ and 3′ ends of all transcripts to further extend our identification of small RNAs in a range of bacteria species. The work has shown that sRNAs are encoded by diverse loci including sequences overlapping mRNAs.
A major focus for the group has been to elucidate the functions of the small RNAs that we and others have identified. Early on, we showed that the OxyS RNA, whose expression is induced in response to oxidative stress, acts to repress translation through limited base pairing with target mRNAs. We discovered that OxyS action is dependent on the Sm–like Hfq protein, which acts as a chaperone to facilitate OxyS RNA–base pairing with its target mRNAs [Reference 1]. Follow-up studies, many in collaboration with the group of Susan Gottesman, have allowed us to learn more about the mechanism by which the Hfq protein facilitates base pairing through multiple RNA–binding domains [Luo X, Zhang A et al., Nucleic Acids Res 2025;in press]. We also have started to explore the role of ProQ, a second RNA chaperone in E. coli, and, by comparing the sRNA–mRNA interactomes by deep sequencing, found that ProQ and Hfq have overlapping as well as competing roles in the cell. It is likely that still other RNA–binding proteins such as KH domain (K homology domain, a conserved nucleic acid recognition motif) proteins are involved in small RNA–mediated regulation [Zamba-Campero M et al., Nat Commun 2024;15:10417].
In addition to characterizing the proteins associated with base-pairing sRNAs, we have been studying the mechanisms by which base pairing alters gene expression. Most characterized interactions between bacterial small RNAs (sRNAs) and their target mRNAs occur near ribosome binding sites, resulting in changes in translation initiation or targeting mRNA decay. However, global RNA–RNA interactome approaches revealed that sRNA base pairing also occurs internal to coding sequences. In a recent study [Thongdee N et al., Mol Cell 2025;85:1824], we examined the impact of sRNA pairing to these internal sequences. Overexpression of the corresponding sRNA led to reduced target-protein levels for two sRNA–mRNA pairs, but there were no differences for five others. By further examining the sRNA–mRNA pairs ArcZ–ligA and ArcZ–hemK, we discovered that ArcZ pairing with the mRNAs leads to translation pausing and increased protein activity. A ligA point mutation that eliminates sRNA pairing resulted in increased sensitivity to DNA damage, revealing the physiological consequences of the regulation. Thus, regulatory RNA pairing in coding sequences can locally slow translation elongation, likely impacting co-translational protein folding and allowing improved incorporation of co-factors or more optimal folding under specific conditions (Figure 1).
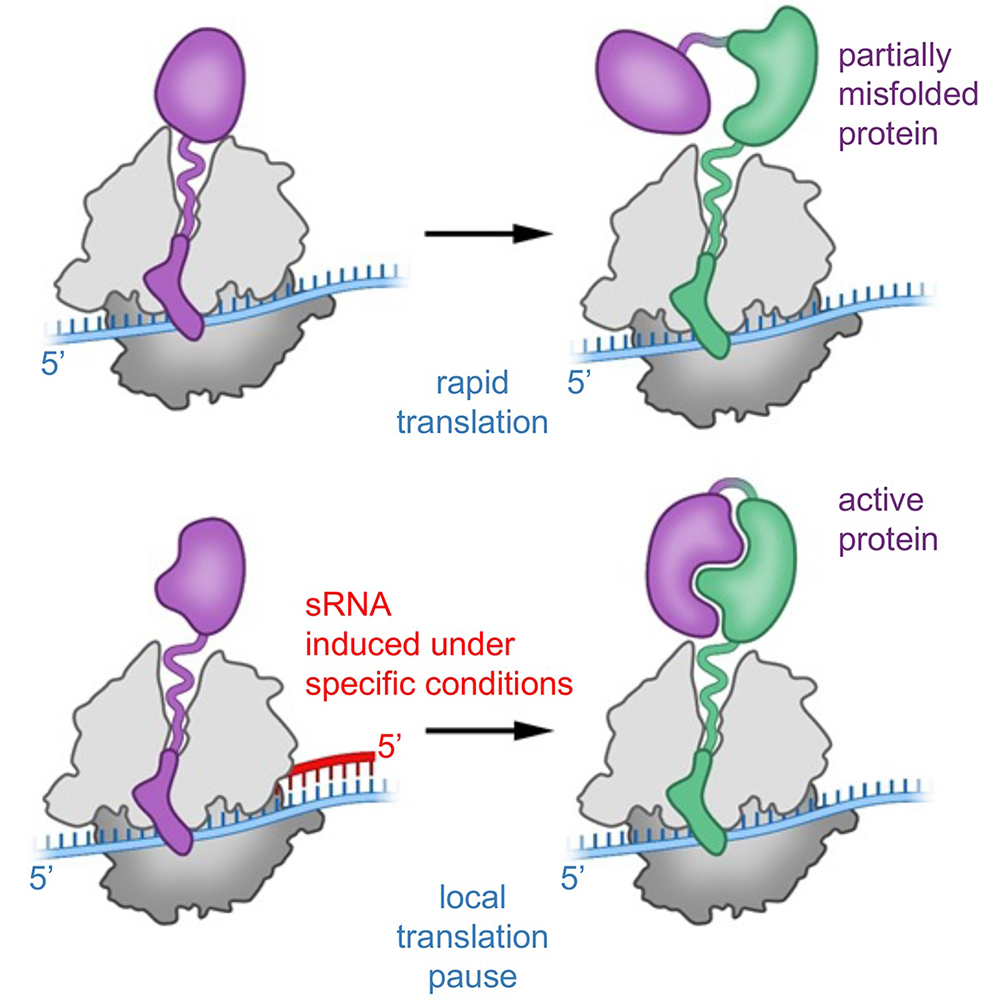
Figure 1. Model of sRNA effects on translation elongation and co-translational protein folding
We suggest that, under certain stress conditions, a non-optimal rate of translation elongation could lead to protein misfolding and reduced activity. Base pairing between an sRNA (induced under the stress conditions) and the mRNA internal to the coding sequence could contribute to the fine-tuning of local translation speed, leading to the corresponding improvements in folding and yield of active protein.
Figure 1. Model of sRNA effects on translation elongation and co-translational protein folding
We suggest that, under certain stress conditions, a non-optimal rate of translation elongation could lead to protein misfolding and reduced activity. Base pairing between an sRNA (induced under the stress conditions) and the mRNA internal to the coding sequence could contribute to the fine-tuning of local translation speed, leading to the corresponding improvements in folding and yield of active protein.
Hfq–binding small RNAs, which act through limited base pairing, are integral to many different stress responses in E. coli and other bacteria as well as during the interaction between bacteria and bacteriophage. Studies of these Hfq–binding sRNAs has given insights into the nuanced control of the regulatory networks as well as into bacterial physiology in general [Papenfort K, Storz G, Cell Chem Biol 2024;31:1571]. For example, we showed that the Spot 42 RNA, whose levels are highest when glucose is present, plays a broad role in catabolite repression by directly repressing genes involved in central and secondary metabolism, redox balancing, and the consumption of diverse non-preferred carbon sources. Similarly, we found that a small RNA derived from the 3′ UTR (untranslated region) of the glnA gene, which encodes glutamine synthetase, impacts E. coli growth under low nitrogen conditions by modulating the expression of genes that affect carbon and nitrogen flux. We also described four UTR–derived sRNAs (UhpU, MotR, FliX, and FlgO), whose expression is controlled by the flagella sigma factor σ28 (fliA) and which have varied effects on flagellin protein levels, flagella number, and cell motility. Intriguingly, MotR, corresponding to the 5′ UTR of an early gene in the flagella regulon, activates flagellar synthesis, while FliX, corresponding to a late gene in the flagella regulon, downregulates flagellar synthesis, illustrating how sRNA–mediated regulation can overlay a complex network enabling temporal control. As more and more sRNAs encoded by 5′ or 3′ UTRs or internal to coding sequences are being found, our observations raise the possibility that phenotypes currently attributed to protein defects are due to deficiencies in unappreciated regulatory RNAs.
One interesting recent observation is that some small RNAs have dual functions in that they act by base pairing but additionally encode a small, regulatory protein. For example, we discovered the Spot 42 RNA also encodes a 15–amino acid protein (denoted SpfP). Overexpression of just the small protein from a Spot 42 derivative deficient in base-pairing activity, or just the base pairing activity from a Spot 42 derivative with a stop codon mutation, both prevented growth on galactose, revealing that the small protein and the small RNA impact the same pathway. As a second example, we found a 164–nucleotide RNA previously shown to encode a 28–amino acid protein (denoted AzuC) also base pairs with the cadA and galE mRNAs to block expression. Interestingly, AzuC translation interferes with the observed repression of cadA and galE by the RNA, and base pairing interferes with AzuC translation, demonstrating that the translation and base-pairing functions compete. We hypothesize that many more dual-function RNAs remain to be discovered and suggest that they can be exploited to control gene expression at multiple levels.
In addition to small RNAs that act via limited base pairing, we have been interested in regulatory RNAs that act by other mechanisms. For instance, early work showed that the 6S RNA binds to and modulates RNA polymerase by mimicking the structure of an open promoter. In another study, we discovered that a broadly conserved RNA structure motif, the yybP–ykoY motif, found in the 5′ UTR of the mntP gene encoding a manganese exporter directly binds manganese, resulting in a conformation that liberates the ribosome-binding site.
Further studies to characterize other Hfq– and ProQ–binding RNAs and their physiological roles and evolution as well as regulatory RNAs that act in ways other than base pairing are ongoing.
Identification and characterization of small regulatory proteins
In our genome-wide screens for small RNAs, we found that a number of short RNAs actually encode small proteins, also denoted microproteins. The correct annotation of the smallest proteins is one of the biggest challenges of genome annotation. Further, there is limited evidence that proteins are synthesized from annotated and predicted short open reading frames (ORFs). Although these proteins have largely been missed, the few small proteins that have been studied in detail in bacterial and mammalian cells have been shown to have important functions in regulation, signaling, and cellular defenses [Burton AT et al., Annu Rev Microbiol 2024;78:1-22]. We thus established a project to identify and characterize proteins of less than 50 amino acids.
We first used sequence conservation and ribosome binding site models to predict ORFs encoding small proteins of 16–50 amino acids in the intergenic regions of the model Escherichia coli genome. We tested expression of these predicted, as well as of previously annotated, small proteins by integrating the sequential peptide affinity tag directly upstream of the stop codon on the chromosome and assaying for synthesis using immunoblot assays. This approach confirmed the synthesis of 20 previously annotated and 18 newly discovered proteins of 16–50 amino acids. In collaboration with Julian Langer, we showed that a subset of these proteins can be identified by mass spectrometry, although this approach is particularly challenging for the smallest proteins [Reference 2]. We also carried out a complementary approach based on genome-wide ribosome profiling of ribosomes arrested on start codons to identify many more candidates; the synthesis of 38 of these small proteins again was confirmed by chromosomal tagging. Most recently, a collaboration with the group of Eugene Koonin, revealed that small protein-coding ORFs could be identified by searches for purifying selection in intergenic regions [References 3, 4]. These studies together with the work of others have documented that E. coli synthesize over 200 small proteins.
Many of the initially discovered proteins were predicted to consist of a single transmembrane alpha-helix and were found to be in the inner membrane. Interestingly, despite their diminutive size, small membrane proteins display considerable diversity in topology and insertion pathways. Additionally, systematic assays for the accumulation of tagged versions of the proteins showed that many small proteins accumulate under specific growth conditions or after exposure to stress.
We are using the tagged derivatives and information about synthesis and subcellular localization, along with many of the approaches the group has used to characterize the functions of small regulatory RNAs, to elucidate the functions of the small proteins. The combined approaches are providing insights into small protein actions, which are relevant to all organisms [Burton AT et al., Annu Rev Microbiol 2024;78:1-22]. For example, we discovered the 49–amino acid inner-membrane protein AcrZ, whose synthesis is increased in response to noxious compounds such as antibiotics and oxidizing agents, associates with the inner-membrane AcrB component of the AcrAB–TolC multidrug efflux pump, a member of the resistance-nodulation-division (RND) superfamily. Mutants lacking AcrZ are sensitive to many, but not all, of the antibiotics transported by AcrAB–TolC due to AcrZ effects on the conformation of the AcrB drug-binding pocket. We also found that synthesis of a 42–amino acid protein MntS is repressed by high levels of manganese by the MntR transcription factor. The lack of MntS leads to reduced activities of manganese-dependent enzymes under manganese-poor conditions, while overproduction of MntS leads to very high intracellular manganese and bacteriostasis under manganese-rich conditions. These and other phenotypes led us to propose that MntS modulates intracellular manganese levels by inhibiting the manganese exporter MntP. Additionally, we showed that the 31–amino acid inner membrane protein MgtS, whose synthesis is induced by very low magnesium by the PhoPQ two-component system (a transcriptional regulator that responds to Mg2+ starvation), acts to increase intracellular magnesium levels and maintain cell integrity upon magnesium depletion. Upon development of a functional tagged derivative of MgtS, we found that MgtS interacts with MgtA to increase the levels of this P-type ATPase magnesium transporter under magnesium-limiting conditions. Correspondingly, the effects of MgtS upon magnesium limitation are lost in a mgtA mutant, and MgtA overexpression can suppress the mgtS phenotype. MgtS stabilization of MgtA provides an additional layer of regulation of this tightly controlled magnesium transporter. A collaborative effort to determine the MgtA structure revealed that this P-type ATPase uniquely forms a dimer [Zeinert R et al., Nat Mol Struct Biol 2025;32:1633] and sets the stage for determining the structure of MgtS in complex with MgtA. Resistance-nodulation-division (RND)–family efflux pumps and P-type ATPase transporters are broadly distributed, also in eukaryotes, and we suggest many more members will be found to be regulated by small proteins.
The ribosome profiling used to identify the intergenic-encoded small proteins revealed there is significant translation initiation within larger open reading frames in the E. coli genome. All five E. coli genes encoding Rpn (recombination-promoting nuclease) proteins have such an internal translation site. We showed that the small, highly variable Rpn C-terminal domains (RpnS), which are translated separately from the full-length proteins (RpnL), directly block the activities of the toxic full-length RpnL proteins, constituting a novel toxin-antitoxin system. The crystal structure of RpnAS revealed a dimerization interface-encompassing helix that has four amino acid repeats, whose number varies widely among strains of the same species. Consistent with strong selection for the variation, we documented that plasmid-encoded RpnP2L protects E. coli against certain phages. We propose that intragenic-encoded small proteins that serve regulatory roles remain to be discovered in all organisms.
The ribosome profiling also revealed that some regulatory sRNAs also encode a small protein and are thus dual-function RNAs. We documented the 109–nucleotide Spot 42 RNA, which is one of the best characterized base-pairing sRNAs in E. coli, encodes the 15–amino acid SpfP protein. Overexpression of just the small protein from a Spot 42 derivative deficient in base-pairing activity resulted in the same phenotype as just the base-pairing sRNA, indicating that the protein and sRNA impact the same pathway. Co-purification experiments revealed that SpfP binds the global transcriptional regulator CRP. This binding blocks the ability of CRP to activate specific genes. Thus, the small protein reinforces the feedforward loop regulated by the base-pairing activity of the Spot 42 RNA. Another dual-function RNA was shown to encode the 28–amino acid, amphipathic helix AzuC protein. We discovered the membrane-associated AzuC protein interacts with GlpD, the aerobic glycerol-3-phosphate dehydrogenase, and increases GlpD membrane association and dehydrogenase activity.
Our work, along with related findings by others in eukaryotic cells, supports our hypothesis that small proteins are an overlooked but important class of proteins, which we continue to study.
Additional Funding
- NICHD Career Development Awards
- NICHD Scientific Director's Award 2025-2026
- NIGMS Postdoctoral Research Associate (PRAT) Program
Publications
- Unexpected richness of the bacterial small RNA world. J Mol Biol 2025 437:169045
- Detection and quantitation of small proteins using mass spectrometry. Mol Cell Proteomics 2025 24:101052
- The hidden bacterial microproteome. Mol Cell 2025 85:1024-1041.e6
- De novo origin of numerous microproteins in enterobacteria. Nucleic Acids Res 2025 In press
- An exciting future for microbial molecular biology and physiology. mBio 2025 16:e0069425
Collaborators
- Philip P. Adams, PhD, Laboratory of Bacteriology, NIAID, NIH, Bethesda, MD
- Yuen-Yan Chang, PhD, Division of Molecular and Cellular Biology, NICHD, Bethesda, MD
- Ryan K. Dale, MS, PhD, Bioinformatics and Scientific Programming Core, NICHD, Bethesda, MD
- Caroline Esnault, PhD, Bioinformatics and Scientific Programming Core, NICHD, Bethesda, MD
- Igor Fesenko, PhD, Computational Biology Branch, NLM/NCBI, NIH, Bethesda, MD
- Pedro H. Franco, Department of Proteomics, Max Planck Institute of Biophysics, Frankfurt, Germany
- Susan Gottesman, PhD, Laboratory of Molecular Biology, Center for Cancer Research, NCI, Bethesda, MD
- Aravind Iyer, PhD, Protein and Genome Evolution Research Group, NLM, NIH, Bethesda, MD
- Mollie W. Jewett, PhD, Division of Immunity and Pathogenesis, University of Central Florida College of Medicine, Orlando, FL
- Eugene V. Koonin, PhD, Evolutionary Genomics Research Group, NLM, NIH, Bethesda, MD
- Julian D. Langer, PhD, Department of Proteomics, Max Planck Institute of Biophysics, Frankfurt, Germany
- Jun Liu, PhD, Department of Microbial Pathogenesis, Yale School of Medicine, New Haven, CT
- Doreen Matthies, PhD, Unit on Structural Biology, NICHD, Bethesda, MD
- Kai Papenfort, PhD, Institute of Microbiology, Friedrich-Schiller-Universität, Jena, Germany
- Marina V. Rodnina, PhD, Max Planck Institute for Multidisciplinary Sciences, Göttingen, Germany
- Harutyun Saakyan, PhD, Computational Biology Branch, NLM/NCBI, NIH, Bethesda, MD
- Ekaterina Samatova, PhD, Max Planck Institute for Multidisciplinary Sciences, Göttingen, Germany
- Svetlana A. Shabalina, PhD, Computational Biology Branch, NLM/NCBI, NIH, Bethesda, MD
- Alexander J. Sort, PhD, Section on Membrane Chemical Physics, NICHD, Bethesda, MD
- Henry Zhang, PhD, Bioinformatics and Scientific Programming Core, NICHD, Bethesda, MD
Contact
For more information, email storzg@mail.nih.gov or visit https://www.nichd.nih.gov/research/atNICHD/Investigators/storz.