Organ and Tissue Formation during Development

- Brant M. Weinstein,
PhD, Head, Section on Vertebrate Organogenesis - Miranda Marvel, PhD, Staff Scientist
- Daniel Castranova, MS, Scientific Technician
- Van Pham, BS, Scientific Technician
- Leah Greenspan, PhD, Postdoctoral Fellow
- Kanako Inoue, PhD, Postdoctoral Fellow
- Jian Ming Khor, PhD, Postdoctoral Fellow
- Aurora Kraus, PhD, Postdoctoral Fellow
- Jong Park, PhD, Postdoctoral Fellow
- Kiyohito Taimatsu, PhD, Postdoctoral Fellow
- Vishakha Vishwakarma, PhD, Postdoctoral Fellow
- Megan Detels, BS, Graduate Student
- Nicholas Kynast, BS, Graduate Student
- Charles White, BS, Graduate Student
- Isabella Cisneros, BS, Postbaccalaureate Fellow
- Justin Gutkowski, BS, Postbaccalaureate Fellow
- Patience Moss, BS, Postbaccalaureate Fellow
- Addison Parker, BS, Postbaccalaureate Fellow
- Jean Prosper Santiago, BS, Postbaccalaureate Fellow
- Zoe Zwick, BS, Postbaccalaureate Fellow
The primary focus of the Section on Vertebrate Organogenesis is to understand how the elaborate networks of blood and lymphatic vessels arise during vertebrate organogenesis and how highly vascularized organs and tissues assemble and function. Blood vessels form a complex, highly ramified, closed circulatory loop that supplies every tissue and organ with oxygen and nutrients. Lymphatic vessels form a similarly complex but entirely separate, highly branched, blind-ended network that drains fluid and waste from the interstitial spaces of tissues, returning them to the blood circulation, and lymphatics also play an important role in immune responses. Our studies on the formation of blood and lymphatic vessels are of substantial clinical interest because of the important roles both types of vessels play in pathologies such as cancer and ischemia.
In addition to our work on vessel development, a major secondary focus of the lab is the role of epigenetics during early development, in particular how DNA methylation and other epigenetic mechanisms that “mark” DNA and chromatin help coordinate cell, tissue, and organ specification and differentiation during development and regeneration. We carry out this work using a novel “EpiTag” zebrafish epigenetic reporter line we recently developed to carry out genetic screens for tissue-specific vertebrate epigenetic regulators.
Our work makes extensive use of the zebrafish (Danio rerio), a small tropical freshwater fish that possesses a unique combination of features that make it a particularly attractive comparative model organism. Zebrafish have most of the same organs and tissues as humans, and they share most of the same genes that we use to direct the assembly and function of these organs and tissues. Zebrafish are also genetically tractable vertebrates, with externally developing, optically clear embryos and larvae that are readily accessible for microscopic imaging and experimental manipulation. These features permit detailed observation of every cell and tissue in a living animal and simple, rapid screening for defects in these cells and tissues.
Our current work includes developing new tools and methods for zebrafish research, uncovering new insights into the vasculature and vascular-related tissues and organs, and carrying out “genetic analysis of epigenetics” using the fish.
Development of new tools and methods for zebrafish research
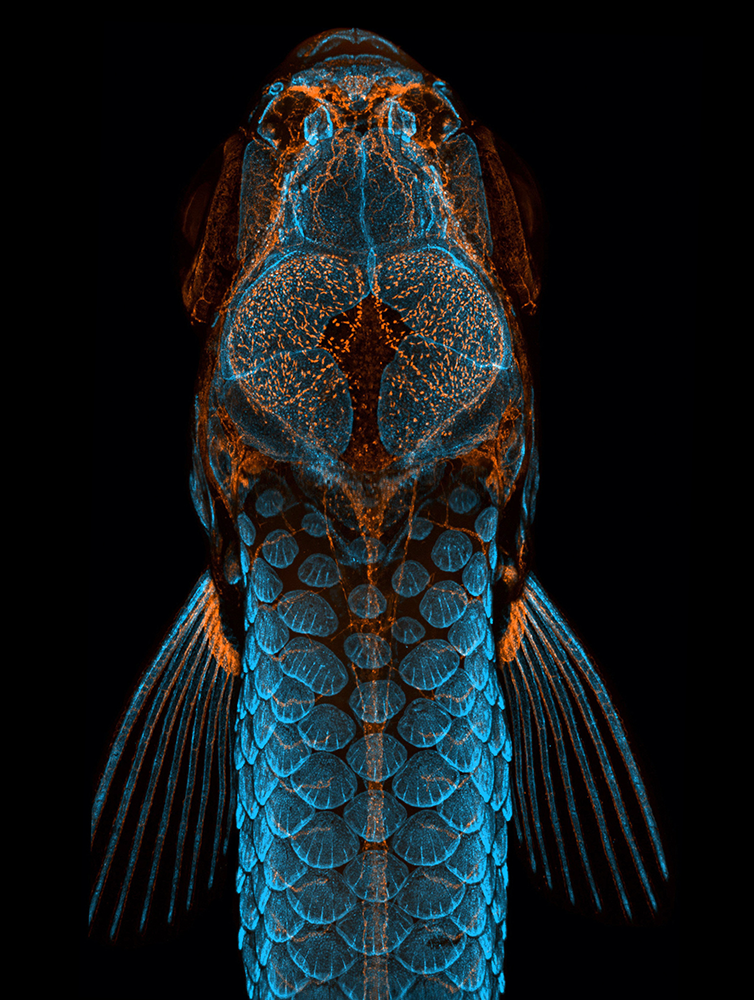
Our laboratory has been a leader in studying vascular development in the zebrafish for nearly three decades, pioneering the use of the zebrafish as a model system for vascular biology. We developed many of the most widely used tools and resources for this research area including confocal microangiography methods [Weinstein BM et al., Nat Med 1995;1:1143-1147], a detailed anatomical atlas of the vasculature [Isogai S et al., Dev Biol 2001;230:278-301], vascular-specific transgenic lines [e.g., Jung HM et al., Development 2017;144:2070], and methods for high-resolution imaging of developing and adult zebrafish [e.g., Jung HM et al., Methods Cell Biol 2016;133:69]. We are also a recognized world leader in sophisticated microscopic imaging, routinely generating award-winning images including a first-place image for the 2020 Nikon Small World competition that was subsequently featured on a 2023 “Life Magnified” United States postage stamp (Figure 1).
We also consistently innovate and share new methods and tools to facilitate zebrafish imaging, including several recently developed methods. These include a method to generate immobilized, pigment-free animals for high-resolution imaging without the need for impractical complex genetic crosses with pigment-defective or “paralyzed” mutants or the use of harmful chemicals to inhibit pigmentation and/or immobilize animals for imaging [Davis AE et al., Zebrafish 2021;18:235]. We also developed a new zebrafish intubation method that permits long-term imaging of anesthetized adult zebrafish for 24 hours or longer with full recovery and survival [Castranova D et al., Development 2022;149:dev199667], a critical tool for carrying out research work using adult zebrafish. We also published a report in 2025 detailing a new tissue clearing method for “super deep imaging” of fluorescently “tagged” zebrafish, which allows for clear, detailed imaging of structures deep within intact animals (Figure 2), even of deep internal regions normally obscured by dense tissues such as cartilage [Reference 1].
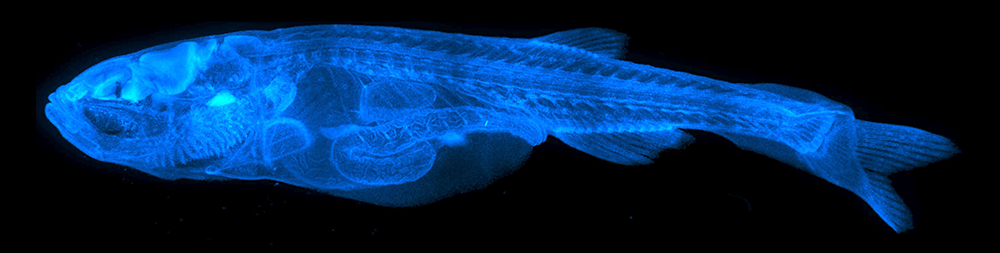
Figure 2. Whole-body “super deep imaging” of an entire 24 day-old zebrafish
Maximum intensity projection confocal image reconstruction of stitched image stacks covering an entire LUCID–cleared, DAPI–stained 24 dpf animal, imaged through a total depth of 655 µm. See Reference 1.
Figure 2. Whole-body “super deep imaging” of an entire 24 day-old zebrafish
Maximum intensity projection confocal image reconstruction of stitched image stacks covering an entire LUCID–cleared, DAPI–stained 24 dpf animal, imaged through a total depth of 655 µm. See Reference 1.
In addition to new tools to facilitate cutting-edge imaging, we have also developed new tools for molecular analysis of zebrafish, including a powerful method for detailed, in-depth “sampling” of genes expressed by specific cells or tissue types in living animals, including the vascular endothelial cells that line the inside of blood and lymphatic vessels [Reference 2]. Unlike methods involving dissociation of organs and tissues into single cells that are then “sorted out” to isolate particular cell types, this new method provides a comprehensive “snapshot” of the genes expressed by specific cell types in their undisturbed, native context. In order to study and better understand the important role that blood and lymphatic vessels play in promoting healing and recovery from the sorts of injuries commonly sustained in human patients, we also recently developed two new methods for modeling skin abrasion wounds [Reference 3] and head trauma [Kraus A et al., bioRxiv 2025;doi: 10.1101/2025.11.14.688311;preprint] in the zebrafish. We describe these new models further in the following section.
New insights into the vasculature and vascular-related tissues and organs
We are using the zebrafish to understand the cellular and molecular mechanisms responsible for proper vessel morphogenesis, for the generation and maintenance of vascular integrity, and for proper repair of the vasculature after injury. We are also using the zebrafish to study previously uncharacterized vascular and vascular-enriched tissues and organs, particularly those with strong human disease relevance.
Genetic analysis of the vasculature
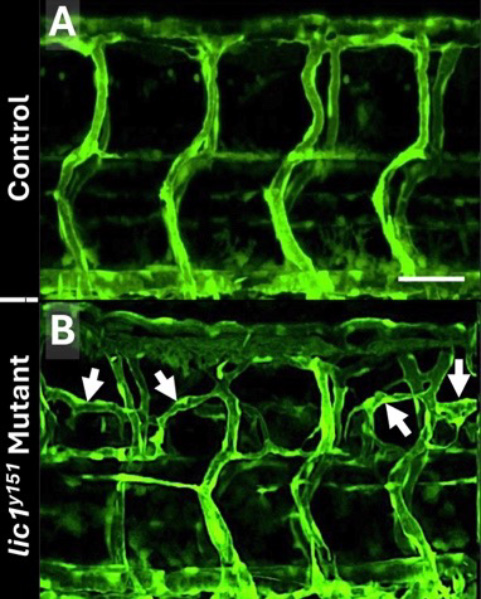
Over the years, we have used genetic screens in the zebrafish to identify mutants with defects in vascular growth, patterning, or integrity and have then characterized the defective genes in these mutants to uncover important new insights into genes and genetic pathways important for vessel growth and vascular-associated disease in humans [e.g., Pillay LM et al., Angiogenesis 2022;25:411]. Recently, we showed that a mutant we identified with vascular overgrowth (Figure 3) was caused by a defect in a gene required for intracellular transport of key pro-angiogenic cell surface receptors [Reference 4], highlighting the important role of intracellular trafficking in controlling receptor availability and resulting angiogenic potential. We also completed another study in 2025 characterizing the function of the human lymphatic disease gene Shoc2, showing that the function of this gene is absolutely required for the development of lymphatic vessels in the zebrafish and identifying a molecular pathway responsible for its activity in the lymphatic endothelial cells [Wilson P et al., bioRxiv 2025;doi: 10.1101/2025.03.26.645567:preprint].
The role of the vasculature in wound healing
As noted above, we developed two new zebrafish models for studying how vessels contribute to healing from skin abrasion wounds and meningeal cerebrovascular injury. Two percent of the US population is plagued by poorly healing cutaneous wounds, most often in the elderly and in diabetics. The reasons for poor healing remain unclear, but vascular dysfunction is thought to be a major contributor. We developed a method for generating reproducible, discrete, readily survivable, cutaneous abrasion wounds in adult zebrafish [Greenspan LJ et al., J Invest Dermatol 2024;144:888] that we are currently using to study the cellular and molecular drivers of cutaneous wound healing angiogenesis, and how these go awry in aged or diabetic zebrafish.
Traumatic head injuries represent another major health burden, affecting nearly 1.4 million people in the United States each year. For many head injury patients, prolonged bleeding and inflammation from damaged vessels in the meninges (thin, highly vascularized tissues that surround and protect the brain) cause long-lasting problems. Unlike mammals with thick, opaque skulls, adult zebrafish have a thin, translucent skull (Figure 4), and their mammalian-like meninges can be easily and noninvasively imaged in living animals, as we recently demonstrated [Venero Galanternik M et al., bioRxiv 2025:doi: 10.1101/2025.04.09.646894:preprint]. We established a novel adult zebrafish meningeal cerebrovascular injury (mCVI) model using carefully calibrated sonication to rupture meningeal blood vessels without breaching the skull or causing damage to the underlying brain [Kraus A et al., bioRxiv 2025; https://doi.org/10.1101/2025.11.14.688311:preprint], and we are currently using this powerful new model to observe vascular regrowth and immune responses to mCVI with unprecedented resolution.
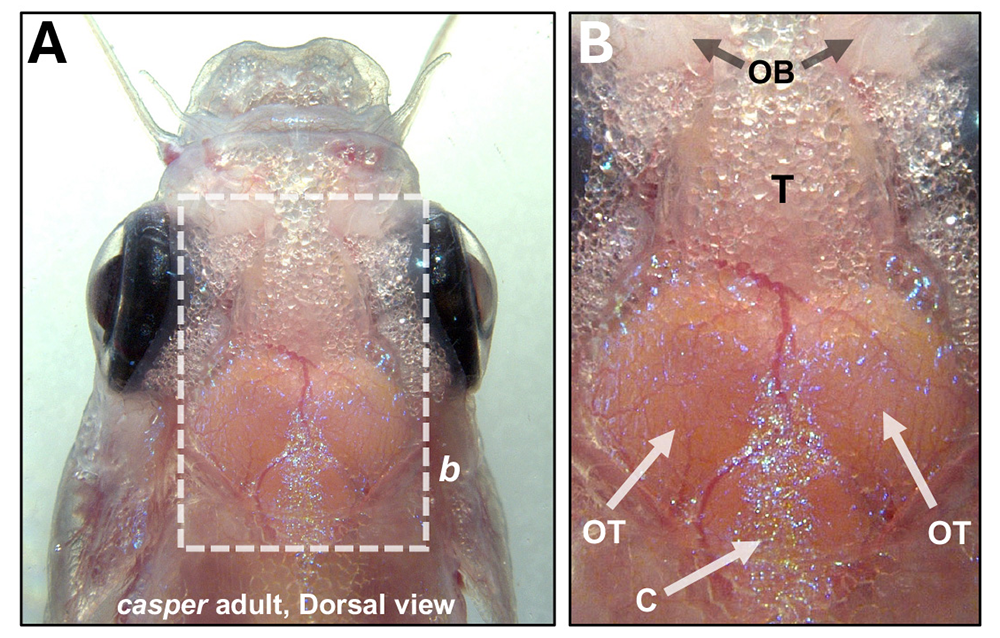
Figure 4. The zebrafish brain can be imaged through its translucent skull.
A. Dorsal view of a pigment-deficient adult casper zebrafish head (anterior up), with the brain surface visible through the thin transparent skull. B. Higher magnification view of the boxed region in panel A, showing olfactory bulbs (OB), telencephalon (T), optic tectum (OT), and cerebellum (C). See Venero Galanternik M et al., bioRxiv 2025.04.09.646894.
Figure 4. The zebrafish brain can be imaged through its translucent skull.
A. Dorsal view of a pigment-deficient adult casper zebrafish head (anterior up), with the brain surface visible through the thin transparent skull. B. Higher magnification view of the boxed region in panel A, showing olfactory bulbs (OB), telencephalon (T), optic tectum (OT), and cerebellum (C). See Venero Galanternik M et al., bioRxiv 2025.04.09.646894.
Vascular-associated tissues and organs
We have maintained a longstanding interest in bringing to light and understanding previously uncharacterized vascular and vascular-enriched tissues and organs in the zebrafish, particularly those with strong human disease relevance. We recently completed a study describing a previously uncharacterized external optically and experimentally accessible immune organ in the zebrafish that we named the Axillary Lymphoid Organ or “ALO” [Reference 5]. Lymph nodes and other secondary lymphoid organs play critical roles in immune surveillance and immune activation in mammals, but their deep internal locations make them challenging to image and study in living animals, and the origins and assembly of these important organs is still not well understood. The ALO is a small translucent organ located on the surface of zebrafish just above the pectoral fin. We showed that the ALO has a structure and cellular composition strongly reminiscent of lymph nodes and other secondary immune organs, making it a superb new model for live imaging of their assembly and function.
We have also been studying the zebrafish gills and their specialized vasculature as a comparative model for the lungs. The high prevalence of lung disease makes it of compelling interest to uncover mechanisms to facilitate lung repair and regeneration, but the relative inaccessibility of mammalian lungs to observation and manipulation has made experimental study challenging. Unlike mammalian lungs, zebrafish gills are readily accessible to live imaging and experimental manipulation, even in adults, and we showed that gills possess many very similar cell types to lungs, including unusual, highly specialized vascular endothelial cells comparable to the so-called “Aerocyte” endothelial cells critical for gas-exchange function in mammalian lung alveoli (Park JS et al., bioRxiv 2025; doi.org/10.64898/2025.11.30.690480). We are currently carrying out additional in-depth studies on these cells with an eye towards translational impact for human lung pathologies.
Genetic analysis of epigenetics
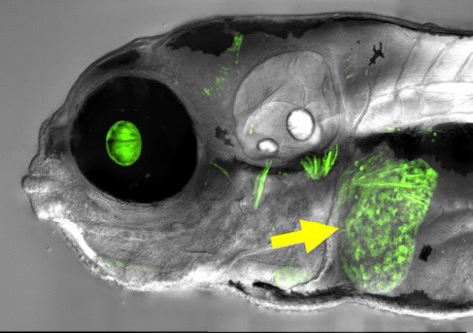
Epigenetic changes (changes in gene expression that occur without altering the DNA sequence) contribute to the generation of cellular diversity during vertebrate development, but the molecular mechanisms involved in organ- and tissue-specific epigenetic regulation in vertebrates such as humans are still relatively unknown. We developed a novel “EpiTag” zebrafish transgenic reporter line that permits real-time visualization of tissue-specific epigenetic silencing or activation in living animals, making it possible for us to monitor dynamic changes in epigenetic regulation in intact animals during development with cellular resolution. Using the EpiTag transgenic line, we performed the first large-scale genetic screen in a vertebrate to identify mutants in regulators of epigenetic gene silencing or activation. Among these mutants we identified a liver-specific epigenetic activation mutant (Figure 5) that provides a new epigenetic model for metabolic-associated fatty liver disease (MAFLD), a disease affecting a quarter of the world’s population for which there is currently no approved drug therapy [Marvel M et al., bioRxiv, 2025;doi: 10.1101/2025.04.21.649821:preprint]. Our research could help point the way towards new epigenetic targets for the treatment of fatty liver disease.
Epigenetic reprogramming is also pivotal for kickstarting dormant developmental programs during regeneration. However, the regulatory mechanisms orchestrating this phenomenon remain elusive. The zebrafish, recognized for its regenerative capabilities, serves as an ideal model to explore this at the molecular level. In addition to serving as a superb tool for studying epigenetic programming occurring during development, and for carrying out genetic screens to identify new epigenetic regulators as described above, the “EpiTag” transgenic line also becomes highly activated in regenerating tissues and organs. This includes very early and striking activation of the EpiTag reporter in regenerating cells following fin amputation (Figure 6), with GFP activation serving as a highly specific marker for cells undergoing epigenetic reprogramming at otherwise inaccessible stages of regeneration. We used EpiTag expression to selectively enrich early-stage regenerating cells and employed a multi-omics approach including ATAC-Seq, bisulfite-Seq, bulk RNA-Seq, and single-cell RNA-Seq to comprehensively profile chromatin accessibility, DNA methylation patterns, gene expression profiles, and single-cell transcriptomes in regenerating cells [Khor JM et al., bioRxiv 2025;2025.04.21.649771]. This approach has led to the identification of novel epigenetic regulatory genes, molecular complexes, and pathways, which we are currently further studying.
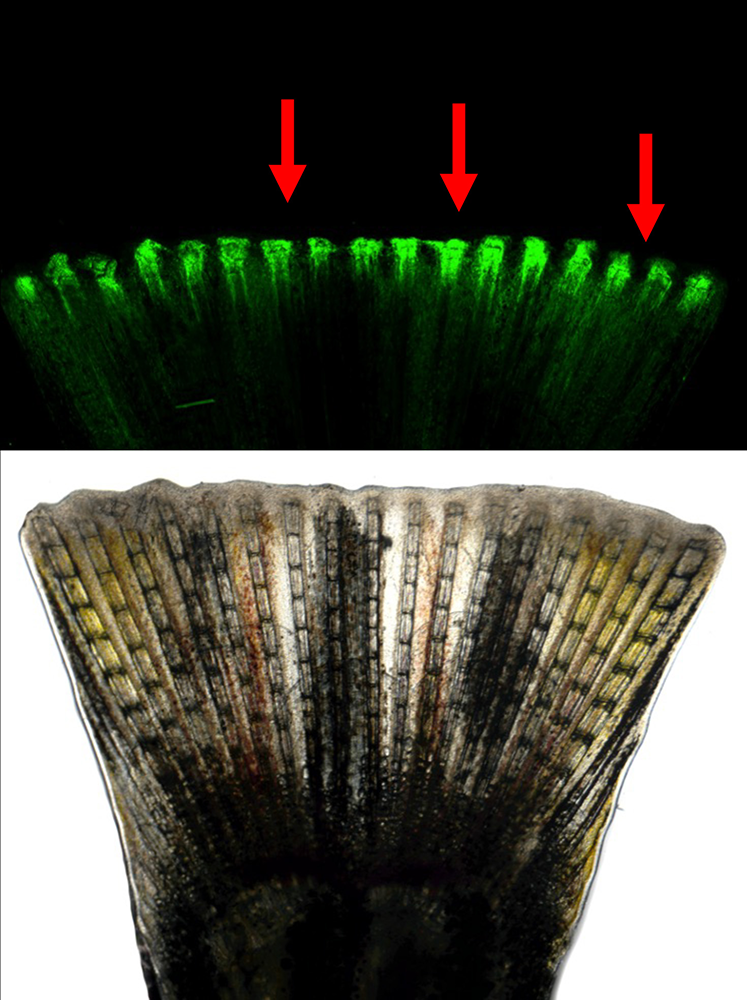
Figure 6. “EpiTag” activation in the regenerating zebrafish fin
Green fluorescence (top) and transmitted light (bottom) imaging of a regenerating EpiTag zebrafish fin one day after amputation. EpiTag fluorescent GFPd2 expression is strongly activated in regenerating cells at the growing fin margin (red arrows). See Khor JM et al., bioRxiv, 2025; 10.1101/2025.04.21.649771.
Figure 6. “EpiTag” activation in the regenerating zebrafish fin
Green fluorescence (top) and transmitted light (bottom) imaging of a regenerating EpiTag zebrafish fin one day after amputation. EpiTag fluorescent GFPd2 expression is strongly activated in regenerating cells at the growing fin margin (red arrows). See Khor JM et al., bioRxiv, 2025; 10.1101/2025.04.21.649771.
Additional Funding
- K99/R00 Pathway to Independence Award (to Leah Greenspan)
- NIGMS Postdoctoral Research Associate Training (PRAT) Fellowship (to Jong Park)
- NICHD Intramural Research Fellowship (to Jian Ming Khor)
- NICHD Career Development Award (to Miranda Marvel)
- NICHD Early Career Award (to Aurora Kraus)
- NICHD Early Career Award (to Jong Park)
- NICHD Early Career Award (to Vishakha Vishwakarma)
- FARE Award (to Jong Park)
- FARE Award (to Kiyohito Taimatsu)
Publications
- Comprehensive 3D imaging of whole zebrafish using a water-based clearing reagent for hard tissues. Zebrafish 2025 22:65-75
- In vivo profiling of the endothelium using ‘AngioTag’ zebrafish. Angiogenesis 2025 28:40
- Live imaging of cutaneous wound healing in zebrafish. J Invest Dermatol 2024 144:888-897
- Angiogenesis is limited by LIC1‐mediated lysosomal trafficking. Angiogenesis 2024 27:943-962
- The axillary lymphoid organ - an external, experimentally accessible immune organ in the zebrafish. J Exp Med 2025 222:e20241435
Collaborators
- Andreas Baxevanis, PhD, Computational and Statistical Genomics Branch, NHGRI, Bethesda, MD
- Harold Burgess, PhD, Section on Behavioral Neurogenetics, NICHD, Bethesda, MD
- Ryan Dale, MS, PhD, Bioinformatics and Scientific Programming Core, NICHD, Bethesda, MD
- Louis Dye, BS, Microscopy and Imaging Core, NICHD, Bethesda, MD
- Jeffrey A. Farrell, PhD, Unit on Cell Specification and Differentiation, NICHD, Bethesda, MD
- Fabio Faucz, PhD, Molecular Genomics Core, NICHD, Bethesda, MD
- Edan Foley, PhD, University of Alberta, Edmonton, CA
- John K. Frazer, PhD, University of Oklahoma Health Sciences Center, Oklahoma City, OK
- Emilia Galperin, PhD, University of Kentucky, Louisville, KY
- Kendrick L. Highsmith, MS, Division of Developmental Biology, NICHD, Bethesda, MD
- Anna Huttenlocher, PhD, University of Wisconsin-Madison, Madison, WI
- James Iben, PhD, Molecular Genomics Core, NICHD, Bethesda, MD
- Sumio Isogai, PhD, Iwate Medical University, Morioka, Japan
- Hyun Min Jung, PhD, University of Illinois, Chicago, IL
- Gennady Margolin, PhD, Bioinformatics and Scientific Programming Core, NICHD, Bethesda, MD
- Yoh-suke Mukouyama, PhD, Laboratory of Stem Cell and Neuro-Vascular Biology, NHLBI, Bethesda, MD
- Jeffrey Mumm, PhD, Johns Hopkins School of Medicine, Baltimore, MD
- Lynn R. Parenti, PhD, Division of Fishes, Department of Vertebrate Zoology, National Museum of Natural History, Smithsonian Institution, Washington, DC
- Lisa M. Price, PhD, Division of Developmental Biology, NICHD, Bethesda, MD
- Eric Raabe, PhD, Johns Hopkins School of Medicine, Baltimore, MD
- Radu V. Stan, MD, PhD, Geisel School of Medicine at Dartmouth, Lebanon, NH
- Amber N. Stratman, PhD, Washington University, St. Louis, MO
- Marina Venero Galanternik, PhD, University of Utah, Salt Lake City, UT
Contact
For more information, email weinsteb@mail.nih.gov or visit https://www.nichd.nih.gov/research/atNICHD/Investigators/weinstein.