You are here: Home > Section on Chromatin and Gene Expression
Chromatin Remodeling and Gene Activation

- David J. Clark, PhD, Head, Section on Chromatin and Gene Expression
- Hope A. Cole, PhD, Postdoctoral Fellow
- Peter Eriksson, PhD, Research Fellow
- Dwaipayan Ganguli, PhD, Visiting Fellow
- Nagarajavel Vivekananthan, PhD, Visiting Fellow
Gene activation involves the regulated recruitment of factors to a promoter in response to appropriate signals, ultimately resulting in the formation of an initiation complex with RNA polymerase II and then transcription. These events must occur in the presence of nucleosomes, which are compact structures capable of blocking transcription at every step. To circumvent the chromatin block, eukaryotic cells possess chromatin-remodeling and nucleosome-modifying complexes. The former (e.g., SWI/SNF complexes) use ATP to drive conformational changes in nucleosomes and to slide nucleosomes along DNA. The latter contain enzymatic activities (e.g., histone acetylases) that modify the histones post-translationally to mark them for recognition by other complexes. Geneticists have described many interesting connections between chromatin components and transcription, but they have lacked a system to investigate the structural basis of the connections. We have developed such a model system, involving native plasmid chromatin purified from the yeast Saccharomyces cerevisiae, to perform high-resolution studies of the chromatin structures of active and inactive genes. Our work reveals that, remarkably, activation correlates with large-scale movements of nucleosomes and conformational changes within nucleosomes over entire genes. Our current work involves two major projects: in the first, we are taking advantage of the recent technological breakthrough in DNA sequencing (high-throughput sequencing) to determine whether our observations concerning gene activation in chromatin can be extrapolated to the entire yeast genome; in the second, we are addressing the role of a sequence-specific DNA-binding histone acetylase, Spt10p, in the cell cycle–dependent regulation of the histone genes.
Transcriptional activation and SWI-SNF-dependent nucleosome mobilization
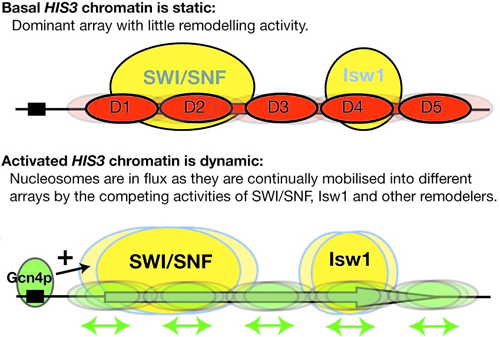
Figure 1. Working model for activation of the HIS3 gene
Basally expressed HIS3 exists in a relatively static chromatin structure approximated by a single ordered nucleosomal array (D1-D5). Activation by Gcn4p initiates nucleosome mobilization by the SWI/SNF and Isw1 complexes. It is suggested that this creates a dynamic chromatin structure in which nucleosomes are shunted back and forth, from one array to another. This would render the DNA transparent to transcription initiation and elongation factors. Not shown: In activated chromatin, nucleosome structure is also remodeled in a SWI/SNF– and Isw1–dependent process.
This work was carried out in collaboration with Bruce Howard. We chose budding yeast as a model organism because biochemical studies of chromatin structure could be combined with molecular genetics. Current models for the role of the SWI/SNF ATP-dependent chromatin remodeling complex in gene regulation are focused on promoters, where the most obvious changes in chromatin structure occur. However, using our plasmid model system with HIS3, a SWI/SNF-regulated gene, we discovered that transcriptional activation creates a domain of remodeled chromatin structure that extends far beyond the promoter, to include the entire gene. We addressed the effects of transcriptional activation on the chromatin structure of HIS3 by mapping the precise positions of nucleosomes in non-induced and transcriptionally activated chromatin. In the absence of the Gcn4 activator, the HIS3 gene is organized into a dominant nucleosomal array. In wild-type chromatin, this array is disrupted, and several alternative, overlapping, nucleosomal arrays are formed. Disruption of the dominant array also requires the SWI/SNF remodeling machine, indicating that the SWI/SNF complex plays an important role in nucleosome mobilization. We propose that Gcn4 stimulates nucleosome mobilization over the entire HIS3 gene by the SWI/SNF complex. We suggest that the net effect of interplay among remodeling machines at HIS3 is to create a highly dynamic chromatin structure. Our work on HIS3 and our earlier work on CUP1 indicate that, at least for these two genes, the target of remodeling complexes is a domain rather than just the promoter. This is an important finding, because it suggests that remodeling complexes act on chromatin domains. A working model is depicted in Figure 1. We speculate that remodeling entire genes might facilitate elongation through nucleosomes by RNA polymerase II.
The advent of high-throughput sequencing has allowed us to analyze nucleosome positioning on a genome-wide scale, confirming and extending our previous observations for HIS3. We are using paired-end sequencing, which increases the accuracy of the position measurements and facilitates the bioinformatics analysis. We have evidence for major disruptions in the chromatin structure of genes when they are activated for transcription. We are exploring the roles of remodeling complexes capable of nucleosome mobilization in disruption of chromatin structure. Our collaborator, Bruce Howard, has written programs to analyze these data. A review on nucleosome positioning has recently appeared (2).
A nucleosomal barrier to transcription by RNA polymerase II in vitro and in vivo
This work was carried out in collaboration with the Studitsky lab. Many eukaryotic genes are regulated at the level of transcript elongation. Nucleosomes are likely targets for this regulation. Previously, the Studitsky Lab showed that nucleosomes formed on very strong positioning sequences (601 and 603) present a high, orientation-dependent barrier to transcription by RNA polymerase II in vitro. The existence of this polar barrier correlates with the interaction of a 16-bp polar barrier signal (PBS) with the promoter-distal histone H3-H4 dimer. In a collaborative study, we found that the polar barrier is relieved by ISW2, an ATP-dependent chromatin remodeler, which translocates the nucleosome over a short distance, such that the PBS no longer interacts with the distal H3-H4 dimer, although it remains within the nucleosome. In vivo, insertion of the 603 positioning sequence into the yeast CUP1 gene results in a modest reduction in transcription, although this reduction is independent of orientation, indicating that the polar barrier can be circumvented. However, the 603-nucleosome is present at the expected position in only a small fraction of cells. Thus, the polar barrier is probably non-functional in vivo because the nucleosome is not positioned appropriately, presumably due to nucleosome sliding activities. We suggest that interactions between polar barrier signals and chromatin remodelers might have significant regulatory potential. A manuscript describing this work is under revision.
Spt10 and Swi4 control the timing of histone H2A/H2B gene activation in budding yeast
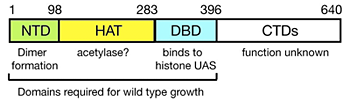
Figure 2. Domain structure of Spt10p
NTD = N-terminal domain; HAT = histone acetylase; DBD = DNA-binding domain; CTDs = C-terminal domains; UAS = upstream activating sequences.
The yeast SPT10 gene encodes a putative histone acetylase (HAT) and has been implicated as a global regulator of core promoter activity. We confirmed that Spt10 has global effects on transcription by expression microarray analysis. Surprisingly, we were able to show that the global effects of Spt10 are the indirect result of defects in chromatin structure, reflecting Spt10's role as activator of the core histone genes. We demonstrated that Spt10 binds specifically and highly cooperatively to pairs of upstream activating sequences (UAS elements) in the core histone promoters (consensus: [G/A]TTCCN6TTCNC), consistent with a direct role in histone gene regulation. No other high-affinity sites are predicted in the yeast genome. Thus, Spt10 is a sequence-specific activator of the histone genes, possessing a DNA-binding domain fused to a likely HAT domain, rather than to a classical activation domain. The DNA-binding domain of Spt10 contains an unusual zinc finger with homology to foamy retrovirus integrase, which we propose to be a sequence-specific DNA-binding protein. Spt10 is a very unusual transactivator, in which the HAT domain, normally recruited as a co-activator to promoters through an activation domain, is attached directly to a sequence-specific DNA-binding domain (Figure 2).
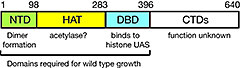
Figure 3. Regulation of the HTA1-HTB1 locus
A Spt10p dimer binds at each pair of UAS elements and activates transcription when a cell cycle–regulated signal is received (which could be Spt21p, or phosphorylation by a cell cycle–dependent kinase). Activation requires SWI/SNF and is repressed by the Hir complex; both effects require the NEG region (blue bar), which contains CCR' (an Abf1p site) and part of the NEG element (black box), which might bind to another protein. (click image to enlarge)
We are now exploring the role of Spt10 in the cell cycle-dependent regulation of the histone genes that is necessary to provide histones for nucleosome assembly during DNA replication. In budding yeast, histones H2A and H2B are expressed from divergent promoters at the HTA1-HTB1 and HTA2-HTB2 loci. We show that Spt10 binds to two pairs of UAS elements in the HTA1-HTB1 promoter: UAS1/UAS2 drive HTA1 expression and UAS3/UAS4 drive HTB1. UAS3 and UAS4 also contain binding sites for the cell cycle regulator SBF (a Swi4-Swi6 heterodimer), which overlap the Spt10 binding sites. Spt10 and SBF binding to UAS3 and UAS4 is mutually exclusive in vitro. Both SBF and Spt10 are bound in cells arrested with alpha-factor, apparently awaiting a signal to activate transcription. Soon after removal of alpha-factor, SBF initiates a small, early peak of HTA1 and HTB1 transcription, which is followed by a much larger peak due to Spt10. Both activators dissociate from the HTA1-HTB1 promoter after expression has been activated. Thus, SBF and Spt10 cooperate to control the timing of HTA1-HTB1 expression. A manuscript describing this work is in revision.
Our current work focuses on the identification of negative regulatory proteins that bind to the histone promoters, which might counteract activation by Spt10p (Figure 3). We have identified candidates and are currently validating them.
Publications
- Clark DJ, Leblanc B. Analysis of DNA supercoiling induced by DNA-protein interactions. Meth Mol Biol 2009;543:523-535.
- Clark DJ. Nucleosome positioning, nucleosome spacing and the nucleosome code. J Biomol Struc Dyn 2010;27:781-793.
Collaborators
- Vasily M. Studitsky, PhD, University of Medicine and Dentistry of New Jersey, Piscataway, NJ
- Bruce H. Howard, MD, Program on Genomics of Differentiation, NICHD, Bethesda, MD
- Blaine Bartholomew, PhD, Southern Illinois University, Carbondale, IL
Contact
For more information, email clarkda@mail.nih.gov or visit ucge.nichd.nih.gov.


