You are here: Home > Section on Chromatin and Gene Expression
Chromatin Remodeling and Gene Activation

- David J. Clark, PhD, Head, Section on Chromatin and Gene Expression
- Peter Eriksson, PhD, Research Fellow
- Tara L. Burke, PhD, Postdoctoral Fellow
- Hope A. Cole, PhD, Postdoctoral Fellow
- Dwaipayan Ganguli, PhD, Visiting Fellow
- Josefina Ocampo, PhD, Visiting Fellow
- Nagarajavel Vivekananthan, PhD, Visiting Fellow
Our main objective is to understand how genes are activated for transcription in the context of chromatin structure. Chromatin is not only a packaging system for DNA in eukaryotic cells but also participates in gene regulation. Many single-gene studies, including our own, have led to various models describing the events that occur in chromatin when genes are activated, both in yeast and higher organisms. But the extent to which such models can be generalized has been unclear. However, the recent advent of high-throughput technologies has led to a plethora of studies addressing the genome-wide locations of sequence-specific DNA-binding proteins, chromatin-remodeling complexes, and nucleosomes. We are exploiting the new technology to map nucleosomes genome-wide in budding yeast and to ask what happens to nucleosomes when genes are activated. We find that many nucleosomes are apparently displaced and that those that remain are re-positioned. We are extending these studies to ask about the roles of transcription activator, ATP-dependent nucleosome remodeling complexes, histone chaperones, and histone modifications in determining the fate of nucleosomes when genes are transcribed. Currently, we are focusing on the roles of the SWI/SNF class of chromatin remodeling machines, which are mutated in various cancers. Our aim is to describe the molecular mechanisms involved in chromatin disruption when genes are activated.
Our second objective is to continue our study of the histone genes in yeast, as a model for the regulation of cell cycle–dependent genes. The histone genes are subject to both positive and negative regulation. We studied the positive regulatory system in depth, demonstrating that Spt10 and SBF are the key positive regulators of the histone genes during the cell cycle. We are now testing a model for histone gene regulation that we proposed recently, which posits a critical role for histone chaperones as part of a negative feedback control system. We are also studying the changes in chromatin structure that occur during the cell cycle, genome-wide, with particular emphasis on the histone gene loci. Thus, we aim to understand the dynamics of chromatin structure during the cell cycle and the roles of trans-activators in this process.
Transcriptional activation and ATP–dependent chromatin remodeling machines
Click image to enlarge.
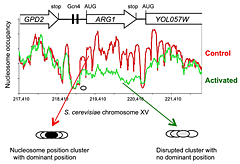
Figure 1.
Activation of the yeast ARG1 gene with 3-aminotriazole results in dramatic loss of nucleosome occupancy and disruption of nucleosome positioning over the coding region, which extends into the downstream YOL057W gene.
The new high-throughput sequencing technology has made it possible to map nucleosomes genome-wide in a relatively simple experiment involving digestion of nuclei with micrococcal nuclease (MNase). Recently, we described a genome-wide paired-end sequencing study of nucleosomes derived from cells treated with 3-aminotriazole (3AT), which induces Gcn4–dependent genes, and from control cells. We concluded that each nucleosome may adopt one of several alternative positions within a cluster and that this cluster organization is genome-wide. The implication is that different cells within a population have different chromatin structures. Thus, binding sites for regulatory factors might be nucleosomal in some cells and in the linker in other cells. Induction with 3AT results in a major disruption of nucleosome positioning, sometimes with altered nucleosome spacing. The most affected genes (50 in all) exhibit a dramatic loss of occupancy over the transcribed region, sometimes extending into neighboring genes. In contrast, nucleosome-depleted promoters are generally unaffected by induction. A small number of genes are repressed by 3AT; these show similar changes in their chromatin structure, but in reverse: loss of nucleosome occupancy is observed in control cells rather than in 3AT–treated cells. Thus, loss of nucleosome occupancy correlates with gene activation (Figure 1).
We examined the extensive nucleosomal rearrangements that occur when genes are activated for transcription by 3AT using a global, bioinformatic, approach (1). At the genomic level, nucleosomes are regularly phased relative to the transcription start site. However, for a subset of strongly induced genes, this phasing is much more irregular after induction, consistent with the loss of some nucleosomes and the re-positioning of the remaining nucleosomes. To address the nature of this rearrangement, we developed the inter-nucleosome distance auto-correlation (DAC) function. At long range, DAC analysis indicates that nucleosomes have an average spacing of 162 bp, as expected. At short range, DAC reveals a 10.25-bp periodicity, implying that nucleosomes in overlapping positions are rotationally related; that is, nucleosome positions overlap by multiples of about 10 bp, the helical period of B-DNA, such that the energy required to bend the DNA around the central core histone octamer is minimally affected. DAC analysis of the 3AT–induced genes suggests that transcription activation coincides with rearrangement of nucleosomes into more irregular arrays. Sequence analysis of the first ("+1") nucleosome on each of the 45 most strongly activated genes reveals a distinctive periodic oscillation in the A/T–dinucleotide occurrence that is present throughout the nucleosome and extends into the linker. This unusual pattern suggests that the +1 nucleosomes might be prone to sliding, thereby facilitating transcription.
We wrote a review of our work, placing it in the context of the chromatin field (2) and described our methodology in detail (3). Currently, we are exploring (i) the roles of remodeling complexes that are capable of nucleosome mobilization, such as SWI/SNF and RSC, in disrupting chromatin structure; and (ii) the structural basis of the chromatin disruption observed on 3AT–induced genes.
Spt10 and SBF control the timing of histone H2A/H2B gene activation in budding yeast.
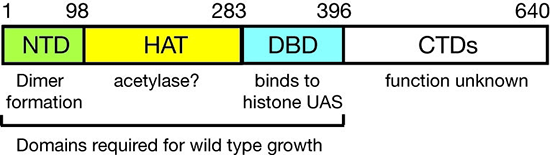
Figure 2. Domain structure of Spt10p
NTD, N-terminal domain; HAT, histone acetylase; DBD, DNA-binding domain; CTDs, C-terminal domains; UAS, upstream activating sequences
We had shown that Spt10 is a very unusual trans-activator, in which a HAT domain, normally recruited as a co-activator to promoters through an activation domain, is attached directly to a sequence-specific DNA–binding domain (Figure 2). More recently, we addressed the role of Spt10 in the cell cycle–dependent regulation of the histone genes, which is necessary to provide histones for nucleosome assembly during DNA replication. Histones H2A and H2B are expressed from divergent promoters at the HTA1–HTB1 and HTA2–HTB2 loci. We showed that Spt10 and the cell-cycle regulator SBF (a Swi4–Swi6 heterodimer) have overlapping binding sites in the HTA1–HTB1 promoter. Both SBF and Spt10 are bound in cells arrested with alpha-factor, apparently awaiting a signal to activate transcription. Soon after removal of alpha-factor, SBF initiates a small, early peak in HTA1 and HTB1 transcription, which is followed by a much larger peak attributable to Spt10. Both activators dissociate from the HTA1–HTB1 promoter after expression has been activated. Thus, SBF and Spt10 cooperate to control the timing of HTA1–HTB1 expression. Our current work has two goals: (i) understanding the dynamics of chromatin structure during the cell cycle at the histone genes and genome-wide; and (ii) testing our proposed model for histone gene regulation (4), which posits that the extent to which histone chaperones are saturated with histones is the critical signal for negative feedback control of transcription (Figure 3).
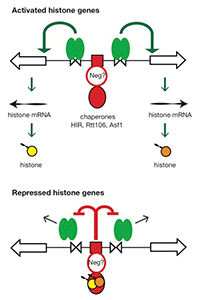
Figure 3. Hypothesis for the regulation of the HTA1-HTB1 locus
Hypothesis: Negative feedback by histones links the positive and negative regulatory (NEG) systems. The histone genes are activated by Spt10 and SBF bound to the histone UAS elements (open triangles), followed by transcription and translation to produce histones. The NEG system inhibits activation through a putative sequence-specific NEG factor (open red circle) bound to the NEG region (red box), which recruits various histone chaperones (HIR, Rtt106, and Asf1; red oval). Inhibition occurs only if histones are bound to the chaperones. The chaperones are fully charged with histone and maximally inhibitory when there is no replicated DNA available in the cell for nucleosome assembly—when replication is complete (outside S-phase) or if replication forks are stalled (e.g., in the presence of hydroxyurea). Outside S-phase, the activators are displaced or degraded (arrows) (4).
Stable RNA polymerase III transcription complexes in vivo
The work described as follows was conducted in collaboration with James Iben, Bruce Howard, and Richard Maraia. TFIIIB and TFIIIC are multi-subunit factors required for transcription by RNA polymerase III. During the course of our nucleosome mapping studies, we discovered that the tRNA genes are occupied by non-nucleosomal complexes, which are in fact stable transcription complexes composed of two multi-subunit factors: TFIIIB and TFIIIC. Accordingly, we described a high-resolution footprint map of TFIIIB–TFIIIC complexes in Saccharomyces cerevisiae, obtained by paired-end sequencing of MNase-resistant DNA (5). On tRNA genes, TFIIIB and TFIIIC form stable complexes with the same distinctive occupancy pattern but in mirror image, termed "bootprints." Global analysis reveals that the TFIIIB–TFIIIC transcription complex exhibits remarkable structural elasticity: tRNA genes vary significantly in length but remain protected by TFIIIC. Introns, when present, are markedly less protected. The RNA polymerase III transcription terminator is flexibly accommodated within the transcription complex and, unexpectedly, plays a major structural role by delimiting its 3′ boundary (Figure 4). The extra TFIIIC (ETC) sites, where TFIIIC binds without TFIIIB, exhibit different bootprints, suggesting that TFIIIC forms complexes involving other factors. We confirmed six ETC sites and found a new site (ETC10). Surprisingly, TFIIIC, but not TFIIIB, interacts with some centromeric nucleosomes, suggesting that interactions between TFIIIC and the centromere may be important in the three-dimensional organization of the nucleus. Currently, we are pursuing this connection between centromeres and transcription complexes.
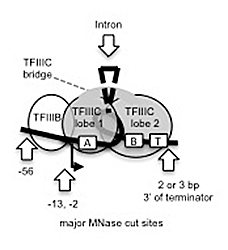
Figure 4. Model for the TFIIIB-TFIIIC complex formed on a tRNA gene
Our global bootprint data are consistent with the coalesced "dumb-bell" structure observed for a TFIIIB-TFIIIC-tRNA gene complex in the electron microscope (described by others). TFIIIC is composed of two lobes connected by a bridge; one lobe binds to the A box, the other to the B box. T: terminator. Red arrows: major sites of micrococcal nuclease (MNase) cleavage.
Additional Funding
- NICHD
Publications
- Cui F, Cole HA, Clark DJ, Zhurkin VB. Transcriptional activation of yeast genes disrupts intragenic nucleosome phasing. Nucl Acids Res 2012;40:10753-10764.
- Cole HA, Nagarajavel V, Clark DJ. Perfect and imperfect nucleosome positioning in yeast. Biochim Biophys Acta 2012;1819:639-643.
- Cole HA, Nagarajavel V, Clark DJ. Genome-wide mapping of nucleosomes in yeast using paired-end sequencing. Methods Enzymol 2012;513:145-168.
- Eriksson PR, Ganguli D, Nagarajavel V, Clark DJ. Regulation of histone gene expression in budding yeast. Genetics 2012;191:7-20.
- Nagarajavel V, Iben JR, Howard BH, Maraia RJ, Clark DJ. Global 'bootprinting' reveals the elastic architecture of the yeast TFIIIB-TFIIIC transcription complex in vivo. Nucl Acids Res 2013;41:8135-8143.
Collaborators
- Bruce H. Howard, MD, Program on Genomics of Differentiation, NICHD, Bethesda, MD
- James Iben, PhD, Program in Genomics of Differentiation, NICHD, Bethesda, MD
- Richard J. Maraia, MD, Program in Genomics of Differentiation, NICHD, Bethesda, MD
- Victor B. Zhurkin, PhD, Laboratory of Cell Biology, Center for Cancer Research, NCI, Bethesda, MD
Contact
For more information, email clarkda@mail.nih.gov or visit ucge.nichd.nih.gov.