You are here: Home > Section on Formation of RNA
Physiological, Biochemical, and Molecular-Genetic Events Governing the Recognition and Resolution of RNA/DNA Hybrids

- Robert J. Crouch, PhD, Head, Section on Formation of RNA
- Susana M. Cerritelli, PhD, Staff Scientist
- Hyongi Chon, PhD, Postdoctoral Fellow
- Lina Gugliotti, PhD, Postdoctoral Fellow
- Kiran Sakhuja, MS, MSc, Research Assistant
- Mariya London, BS, Technical IRTA
Damaged DNA is one of the leading causes of many human diseases and disorders. We study the formation and resolution of RNA/DNA hybrids, which occur during DNA and RNA synthesis. Such hybrid molecules may lead to increased DNA damage but may also play critical roles in normal cellular processes. We are interested in how RNA/DNA hybrids are resolved and in the role that ribonucleases H (RNases H) play in their elimination. Two classes of RNases H are present in most organisms. Our studies have shown that mice deleted for the Rnaseh1 gene arrest embryonic development at day 10 as a result of failure to amplify mitochondrial DNA. Others have found that the Aicardi-Goutières Syndrome (AGS), a severe neurological disorder with symptoms appearing at or soon after birth, can be caused by defective human RNase H2. We employ molecular-genetic and biochemical tools and yeast and mouse models in our research.
RNA/DNA hybrids can be composed of one strand of RNA and a second strand of DNA or can be in the form of an R-loop in which the two DNA strands are separated, with only one hybridized to RNA while the other is single-stranded DNA. A third "hybrid" is a single ribonucleotide in duplex DNA. The first two types of hybrid are substrates for class I and II RNases H. The third is uniquely recognized by type 2 RNases H (Figure 1).
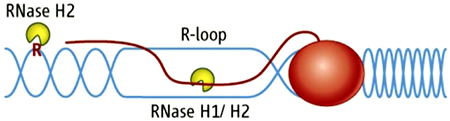
Figure 1. RNA/DNA hybrids and single rNMPs in DNA are substrates for RNase H2.
DNA is in blue, RNA is red line and letter R. The red oval represents RNA polymerase.
Simple (non-R-loop) RNA/DNA hybrids are formed when the HIV-AIDS virus copies its genomic RNA into DNA using a reverse transcriptase (RT). The RT also has an RNase H domain that is structurally and functionally similar to the class I cellular RNase H and is necessary for several steps of viral DNA synthesis. It is known that, during RNA synthesis, R-loops can form and that aberrant R-loop formation may result in chromosome breakage. However, R-loop formation has been observed in the normal recombination process of switching (recombination) from one form of immunoglobulin to another, resulting in different isoforms of antibodies. Single-ribonucleotide hybrids occur during DNA replication, and the ribonucleotide must be removed, a process initiated by RNase H2.
Contrasts between Class I and II RNases H
Previous Annual Reports detailed our work on RNase H1. RNase H1 recognizes the 2′-OH of four consecutive ribonucleotides while the DNA strand is distorted to fit in a pocket of the enzyme. Thus, the enzyme requires more than one ribonucleotide for cleavage of RNA in RNA/DNA hybrids. RNases H1 consist of a single polypeptide in both eukaryotes and prokaryotes. In contrast, RNase H2 is a complex of three different polypeptides in eukaryotes but is a single polypeptide in prokaryotes. One of the subunits of the heterotrimeric RNase H2 of eukaryotes is similar in its primary amino acid sequence to the prokaryotic enzyme. RNase H2 can recognize and cleave a single ribonucleotide or the transition from the ribonucleotide in the case of RNA-primed DNA synthesis (e.g., rrrrrDDDD in DNA—italics indicate transition from ribonucleotide to deoxyribonucleotide—see Figure 1 and references 1–5).
Dual activities of RNase H2 and Aicardi-Goutières syndrome
Eukaryotic RNases H2 recognize and resolve two chemically distinct structures (Figure 1)—the "small" difference being the oxygen present in RNA. The same enzyme and catalytic mechanism of hydrolysis are employed for both types of substrate. The absence of an RNase H2 subunit, mutations that lower amounts of catalytic activity, or failure to properly interact with in vivo substrates could all be deleterious, particularly in multi-cellular organisms. Aicardi-Goutières syndrome (AGS)–related mutations are present in any of the three RNase H2 subunits. Of the 29 different RNase H2 mutations associated with AGS, eight are in the A subunit, 14 in the B subunit, and 7 in the small C subunit (3). We previously expressed in E. coli and purified different human RNases H2 with mutations corresponding to several of those seen in AGS patients' RNases H2; we found only one with significant loss of RNase H2 activity (4). The structure we determined allows us to locate all known mutations in RNase H2 causing AGS (3). The wide distribution of these mutations suggests that modest changes in stability, interaction with other unknown proteins, and loss of catalysis can all cause AGS.
The difference between substrates cleaved by bacterial and eukaryotic RNases H2 in vitro raises the question as to how the eukaryotic enzymes have expanded their repertoire. The interaction of the Tyr residue with the deoxyribose in T. maritima RNase H2 would not be possible were a ribose present owing to a conflict with the 2′-OH of the ribose. One possibility for expanded substrate recognition is that the conserved Tyr is more flexible in eukaryotic RNases H2 than in bacterial enzymes. This question is of importance because single-ribonucleotide substrates appear during DNA replication and repair whereas R-loop structures could occur any time when transcription takes place (not necessarily limited to replication and repair).
To examine the consequences of failure to remove persistent RNA/DNA hybrids and single ribonucleotides joined to DNA in vivo, it was necessary to modify RNase H2 such that only one of these two types of substrate was not cleavable. Based on a rational design comparing the structures of RNase H2 and H3, we unlinked the two activities to yield an enzyme that only cleaves RNA/DNA, leaving single ribonucleoside monophosphates (rNMPs) attached to DNA. RNases H2 and H3 have similar 3D structures, but RNase H3 does not cleave single rNMPs in DNA. We first examined in vitro activities of this enzyme using, among others, the ribonucleotide excision repair (RER) assay recently developed by our collaborators Peter Burgers and Justin Sparks (5). The in vitro results show complete lack of removal of single ribonucleotides but only modest reduction in hydrolysis of RNA/DNA hybrids. In vivo, we found that the signature 2–5 bp deletions in the yeast CAN1 gene, when RNase H2 activity is absent, were also observed when our mutant RNase H2 replaced the wild-type enzyme. The mutant enzyme was able to resolve RNA/DNA hybrids that either RNase H2 or RNase H1 use as in vivo substrates. An important new observation associates RNase H2 RNA/DNA with resolution of a unique substrate—an activity that cannot be replaced by RNase H1—and most likely indicates special access to this class of hybrids via contacts with other cell components. An example of this special class occurs when homologous recombination is defective due to loss of SGS1 helicase function. Thus, the synthetic defect observed in sgs1D, rnh201D strains results from problems associated with persistent R-loops. Given that our RNase H2 mutant enzyme resolves these R-loops, the synthetic defect is absent when the mutant RNase H2 is present in a strain deleted for SGS1.
The RNaseH2A G37S mutation, which is found in a few AGS patients, results in limited activity in vitro (4). We investigated how the corresponding mutation in yeast RNase H2 (G42S) protein functions. In vitro it had very poor activity on RNA/DNA hybrids AND on cleavage of single rNMPs in DNA. In vivo it was almost as defective as a deletion of an RNase H2 subunit for removing single rNMPs (measured by 2–5 bp deletions in the CAN1 gene), had limited activity in removing R-loops accessible to either RNase H2 or H1, yet dislayed remarkably good activity thought to involve interaction with other proteins involved in R-loops associated with DNA replication/repair.
We are currently working with mouse models of AGS to clarify what activity is the one most important for causing AGS.
Mitochondrial DNA and RNase H1 in mouse
In collaboration with Ian Holt, we are examining the roles of RNase H1 in mitochondrial DNA replication by using, among other techniques, analysis of intermediates on two-dimensional gels. Our findings thus far indicate that elevated expression of RNase H1 in mitochondria alters mtDNA replication. In addition, data obtained in Holt's laboratory by John Holmes supporting the existence of RNA/DNA hybrids as replication intermediates indicate a bi-directional mode of DNA replication for this organelle. This work continued last year.
Additional Funding
- Scientific Directors Intramural Award
Publications
- Cerritelli SM, Crouch RJ. Ribonuclease H: the enzymes from eukaryotes. FEBS J 2009;276:1494-1505.
- Cerritelli SM, Chon H, Crouch, RJ. A new twist for topoisomerase. Science 2011;332:1510-1511.
- Figiel M, Chon H, Cerritelli SM, Cybulska M, Crouch RJ, Nowotny M. The structural and biochemical characterization of human RNase H2 complex reveals the molecular basis for substrate recognition and Aicardi-Goutières Syndrome defects. J Biol Chem 2011;286:10540-10550.
- Chon H, Vassilev A, DePamphilis ML, Zhao Y, Zhang J, Burgers PM, Crouch RJ, Cerritelli SM. Contributions of the two accesory subunits, RNASEH2B and RNASEH2C, to the activity and properties of the human RNase H2 complex. Nucleic Acids Res 2009;37:96-110.
- Sparks JL, Chon H, Cerritelli SM, Kunkel TA, Johansson E, Crouch RJ, Burgers PM. RNase H2-initiated ribonucleotide excision repair. Mol Cell 2012;47:980-986.
Collaborators
- Peter Burgers, PhD, Washington University, St. Louis, MO
- Ian Holt, PhD, MRC-Dunn Nutrition Unit, Cambridge, UK
- Herbert C. Morse, MD, Laboratory of Immunopathology, NIAID, Bethesda, MD
- Marcin Nowotny, PhD, International Institute of Molecular and Cell Biology, Warsaw, Poland
- Keiko Ozato, PhD, Program in Genomics of Differentiation, NICHD, Bethesda, MD
- Roger Woodgate, PhD, Laboratory of Genomic Integrity, NICHD, Bethesda, MD
Contact
For more information, email crouch@helix.nih.gov or visit www.nichd.nih.gov/research/atNICHD/Investigators/crouch.

