Protein Sorting in the Endomembrane System

- Juan S. Bonifacino, PhD, Head, Section on Intracellular Protein Trafficking
- Rafael Mattera, PhD, Staff Scientist
- Xiaolin Zhu, Nurse, Technician
- Jing Pu, PhD, Research Fellow
- Raffaella De Pace, PhD, Visiting Fellow
- Saikat Ghosh, PhD, Visiting Fellow
- Carlos M. Guardia, PhD, Visiting Fellow
- Morie Ishida, PhD, Visiting Fellow
- Rui Jia, PhD, Visiting Fellow
- Tal Keren-Kaplan, PhD, Visiting Fellow
- Elodie Mailler, PhD, Visiting Fellow
- Amra Saric, PhD, Visiting Fellow
- Lucas M. Djavaherian, BA, Postbaccalaureate Student
- Gerard Walker, BA, Postbaccalaureate Student
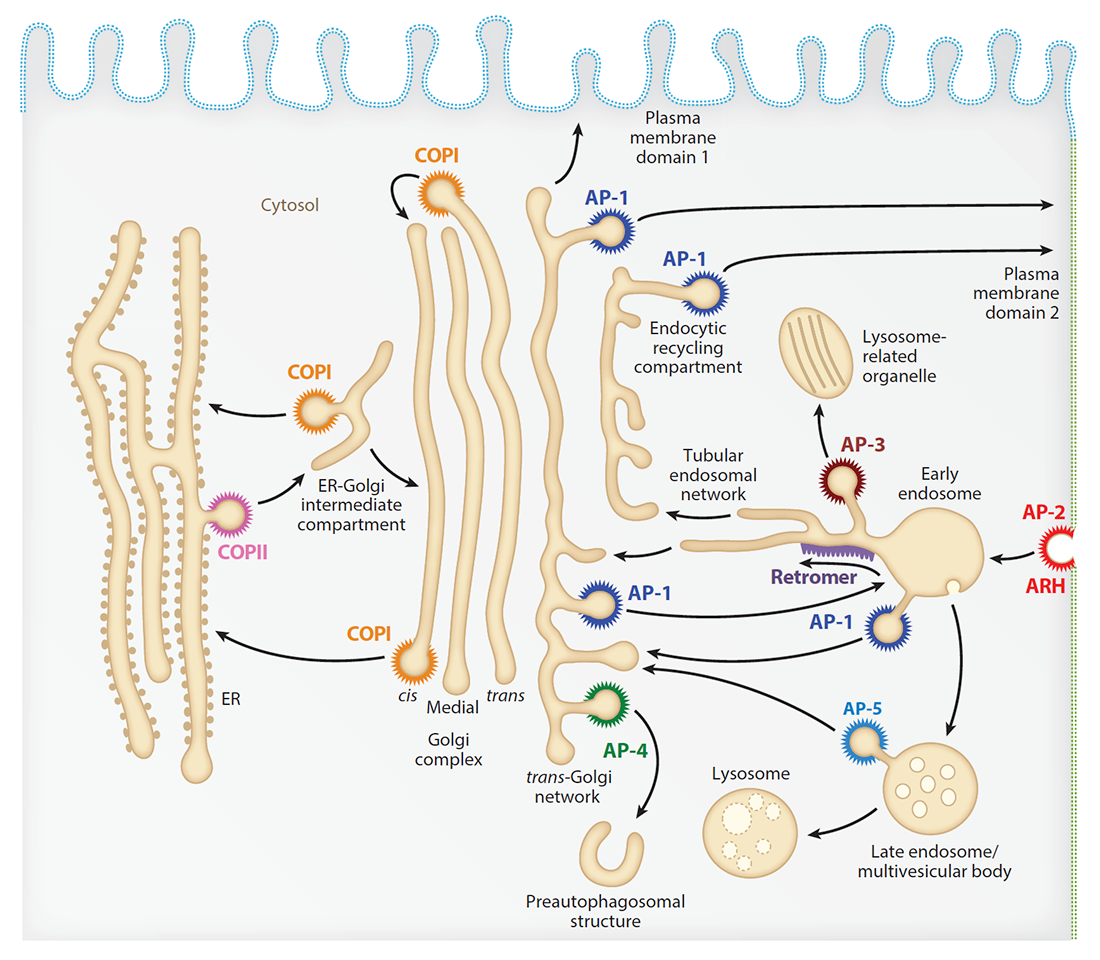
Our laboratory investigates the molecular mechanisms by which transmembrane proteins (referred to as “cargo”) are sorted to different compartments of the endomembrane system in eukaryotic cells. The system comprises an array of membrane-enclosed organelles including the endoplasmic reticulum (ER), the Golgi apparatus, the trans-Golgi network (TGN), endosomes, lysosomes, lysosome-related organelles (LROs, e.g., melanosomes), and various domains of the plasma membrane in polarized cells, such as epithelial cells and neurons (Figure 1). Transport of cargo between these compartments is mediated by vesicular/tubular carriers that bud from a donor compartment, translocate through the cytoplasm, and eventually fuse with an acceptor compartment. Work in our laboratory focuses on the molecular machineries that mediate such processes, including (1) sorting signals and adaptor proteins that select cargo proteins for packaging into the transport carriers, (2) microtubule (MT) motors and organelle adaptors that drive movement of the transport carriers and other organelles through the cytoplasm, and (3) tethering factors that promote fusion of the transport carriers to acceptor compartments. We study the machineries in the context of different intracellular transport pathways, including endocytosis, recycling to the plasma membrane, retrograde transport from endosomes to the TGN, biogenesis of lysosomes and LROs, autophagy, and polarized sorting in epithelial cells and neurons. We apply knowledge gained from this basic research to the elucidation of disease mechanisms, including congenital disorders of protein traffic, such as the pigmentation and bleeding disorder Hermansky-Pudlak syndrome (HPS), hereditary spastic paraplegias (HSPs), and other neurodevelopmental disorders.
Figure 1. Schematic representation of the endomembrane system of eukaryotic cells showing the localization of coats involved in protein sorting
The FHF complex mediates perinuclear distribution of AP-4 and its cargo ATG9A.
Over 20 years ago, we discovered the heterotetrameric adaptor protein complex 4 (AP-4) as a component of a protein coat associated with the TGN. Other groups then showed that mutations in AP-4 subunits cause a complicated form of autosomal-recessive hereditary spastic paraplegia termed AP-4-deficiency syndrome. Recent studies from our lab demonstrated that AP-4 mediates export of the transmembrane autophagy protein ATG9A from the TGN to pre-autophagosomal structures, and that the export contributes to the maintenance of autophagic homeostasis. Failure to export ATG9A from the TGN in AP-4–deficiency patients may underlie the pathogenesis of the syndrome. This past year, we sought to identify additional proteins that cooperate with AP-4 in ATG9A trafficking. Using affinity purification and mass spectrometry, we identified the FHF complex (a protein complex thought to promote vesicle trafficking and/or fusion) as a novel AP-4 accessory factor. Knockdown of FHF subunits resulted in dispersal of AP-4 and ATG9A from the perinuclear region of the cell, consistent with the previously demonstrated role of the FHF complex in coupling organelles to the MT retrograde motor dynein-dynactin. The findings uncovered an additional mechanism for the distribution of ATG9A and provided further evidence for a role of protein coats in coupling transport vesicles to MT motors.
Structure of human ATG9A, the only transmembrane protein of the core autophagy machinery
The identification of ATG9A as an AP-4 cargo prompted us to examine the structure and function of the protein. ATG9A is the only transmembrane component of the core autophagy machinery and contributes to autophagosome biogenesis by mechanisms that are not well understood. In collaboration with the groups of Anirban Banerjee, Jiansen Jiang, and José Faraldo-Gómez, we succeeded in obtaining a 2.9 Å–resolution cryo–EM structure of human ATG9A. The structure revealed a novel fold with a homotrimeric domain-swapped architecture, multiple membrane spans, and a network of branched cavities, consistent with ATG9A being a membrane transporter. In addition, structure-guided molecular simulations predicted that ATG9A causes membrane bending, explaining the localization of the protein to small vesicles and highly curved edges of growing autophagosomes.
Regulation of LC3B levels by ubiquitination and proteasomal degradation
In addition to working on ATG9A structure, we investigated the mechanisms of autophagy regulation. To this end, we conducted a genome-wide CRISPR-Cas9 knockout screen using cells expressing endogenous LC3B (a microtubule-associated protein that is central to the autophagy pathway) tagged with GFP-mCherry as a reporter, an approach that allowed us to identify the ubiquitin-activating enzyme UBA6 and the hybrid ubiquitin-conjugating enzyme/ubiquitin ligase BIRC6 as novel autophagy regulators. We found that the enzymes cooperate to mediate mono-ubiquitination and proteasomal degradation of LC3B, thus limiting the pool of LC3B available for autophagy. Depletion of UBA6 or BIRC6 raised the level of cytosolic LC3B, enhancing the degradation of autophagy adaptors and the clearance of intracellular proteins aggregates. The finding could be the basis for the development of pharmacological inhibitors of UBA6 or BIRC6 for the treatment of protein-aggregation disorders.
ARL8 relieves SKIP autoinhibition to enable coupling of lysosomes to kinesin-1.
The lab also focusses on the mechanisms that drive movement of organelles within the cytoplasm. Long-range movement of organelles relies on coupling to microtubule motors, a process that is often mediated by adaptor proteins. In many cases, the coupling involves organelle- or adaptor-induced activation of the microtubule motors by conformational reversal of an auto-inhibited state. This past year, we discovered that a similar regulatory mechanism operates for an adaptor protein named SKIP (also known as PLEKHM2). SKIP binds to the small GTPase ARL8 on the lysosomal membrane to couple lysosomes to the anterograde microtubule motor kinesin-1. Structure-function analyses of SKIP revealed that the C-terminal region, comprising three PH domains, interacts with the N-terminal region, comprising ARL8- and kinesin-1–binding sites. The interaction inhibits coupling of lysosomes to kinesin-1 and, consequently, lysosome movement toward the cell periphery. We also found that ARL8 not only recruits SKIP to the lysosomal membrane, but also relieves SKIP auto-inhibition, promoting kinesin-1–driven, anterograde lysosome transport. The findings demonstrate that SKIP is not merely a passive connector of lysosome-bound ARL8 to kinesin-1 but is itself subject to both intra- and inter-molecular interactions that regulate its function
Synaptic vesicle precursors and lysosomes are transported by different mechanisms in the axon of mammalian neurons.
BORC is a multi-subunit complex that induces the recruitment of ARL8 to membranes. Previous work showed that BORC promotes coupling of mammalian lysosomes and Caenorhabditis elegans synaptic vesicle precursors (SVPs) to kinesins for anterograde transport of these organelles along microtubule tracks, raising the possibility that axonal lysosomes and SVPs are the same or related organelles. Analysis of wild-type and BORC–knockout mice, however, showed that SVPs and lysosomes are transported separately, and that only lysosomes depend on BORC for axonal transport in mammalian neurons. The findings thus demonstrated that SVPs and lysosomes are distinct organelles that rely on different machineries for axonal transport in mammalian neurons.
The Parkinson’s disease protein LRRK2 interacts with the GARP complex to promote retrograde transport to the TGN.
In an extension of our work on the role of the GARP complex as a TGN–tethering factor, we examined the physical and functional interactions of GARP with the protein leucine-rich repeat kinase 2 (LRRK2), in collaboration with the lab of Mark Cookson. Mutations in LRRK2 cause Parkinson's disease (PD); however, the precise function of LRRK2 remains unclear. We found that LRRK2 interacts with the VPS52 subunit of GARP (Golgi-associated retrograde protein). LRRK2 further interacts with the Golgi SNAREs (proteins that mediate vesicle fusion) VAMP4 and Syntaxin-6 and acts as a scaffold that stabilizes GARP–SNAREs complex formation. LRRK2 thus influences retrograde trafficking pathways in a manner that depends on its GTP–binding and kinase activity, an action that is exaggerated by mutations associated with PD and which can be blocked by kinase inhibitors. Disruption of GARP sensitizes dopamine neurons to mutant LRRK2 toxicity in Caenorhabditis elegans, showing that the pathways are interlinked in vivo.
Publications
- Jia R, Bonifacino JS. Negative regulation of autophagy by UBA6-BIRC6-mediated ubiquitination of LC3. eLife 2019;8:e50034.
- Guardia CM, Tan XF, Lian T, Rana MS, Zhou W, Christenson ET, Lowry AJ, Faraldo-Gómez JD, Bonifacino JS, Jiang J, Banerjee A. Structure of human ATG9A, the only transmembrane protein of the core autophagy machinery. Cell Rep 2020;31:107837.
- Mattera R, Williamson CD, Ren X, Bonifacino JS. The FTS-Hook-FHIP (FHF) complex interacts with AP-4 to mediate perinuclear distribution of AP-4 and its cargo ATG9A. Mol Biol Cell 2020;31:963-979.
- De Pace R, Britt DJ, Mercurio J, Foster AM, Djavaherian L, Hoffmann V, Abebe D, Bonifacino JS. Synaptic vesicle precursors and lysosomes are transported by different mechanisms in the axon of mammalian neurons. Cell Rep 2020;31:107775.
- Keren-Kaplan T, Bonifacino JS. ARL8 relieves SKIP autoinhibition to enable coupling of lysosomes to kinesin-1. Curr Biol 2020;31(3):540-554.E5.
Collaborators
- Anirban Banerjee, PhD, Unit on Structural and Chemical Biology of Membrane Proteins, NICHD, Bethesda, MD
- Mark Cookson, PhD, Laboratory of Neurogenetics, NIA, Bethesda, MD
- José Faraldo-Gómez, PhD, Theoretical Molecular Biophysics Laboratory, NHLBI, Bethesda, MD
- Jiansen Jiang, PhD, Laboratory of Membrane Proteins and Structural Biology, NHLBI, Bethesda, MD
- Yihong Ye, PhD, Laboratory of Molecular Biology, NIDDK, Bethesda, MD
Contact
For more information, email bonifacinoj@helix.nih.gov or visit https://www.nichd.nih.gov/research/atNICHD/Investigators/bonifacino.