Genes and Signals Regulating Mammalian Hematopoiesis

- Paul E. Love, MD, PhD, Head, Section on Cellular and Developmental Biology
- LiQi Li, MD, PhD, Research Fellow
- Amy Palin, PhD, Postdoctoral Fellow
- Claude Warzecha, PhD, Postdoctoral Fellow
- Zhenhu Li, PhD, Visiting Fellow
- Bin Zhao, PhD, Visiting Fellow
- Dalal El-Khoury, BS, Technician
- Jan Lee, BS, Technician
- Miles Oliva, BS, Postbaccalaureate Intramural Research Training Award Fellow
- Daniel Stamos, BS, Postbaccalaureate Intramural Research Training Award Fellow
Our research focuses on the development of the mammalian hematopoietic system. Of particular interest is the characterization of signal-transduction molecules and pathways that regulate T cell maturation in the thymus. Current projects include the generation of transgenic and conditional deletion mutants to evaluate the importance of T cell antigen receptor signaling at specific stages of T cell development. We are also using microarray gene profiling to identify molecules that are important for thymocyte selection, a process that promotes the survival and further development of functional T cells and the death of auto-reactive T cells, thereby preventing autoimmunity.
A newer project involves analyzing the function of Themis, a T cell–specific signaling protein recently identified by our laboratory. Another recently initiated area of investigation focuses on hematopoietic stem cells (HSCs), which give rise to all blood cell lineages. We have begun to characterize the genes that are important for the generation and maintenance of HSCs and for their differentiation into specific hematopoietic cell types. The studies revealed a critical function for one protein (Ldb1) in controlling the self-renewal/differentiation cell-fate decision in both HSCs and erythroblasts by acting as a key component of multi-subunit DNA–binding complexes. Global (ChIP-seq) screening for Ldb1–complex DNA–binding sites identified many targets for Ldb1–mediated regulation of transcription in hematopoietic cells, demonstrating an important role for Ldb1 in hematopoietic gene regulation. Current work on Ldb1 includes an examination of a potential role for this protein in regulating self-renewal of T cell progenitors in the thymus and in the genesis of T cell Acute Lymphoblastic Leukemia (T-ALL), one of the most common childhood malignancies.
T cell antigen receptor signaling in thymocyte development
Much of our research focuses on the role of T cell antigen receptor (TCR) signal transduction in thymocyte development. Signal transduction sequences, termed immunoreceptor tyrosine-based activation motifs or ITAMs, are contained within four distinct subunits of the multimeric TCR complex (CD3-zeta, CD3-gamma, CD3-delta, and CD3-epsilon). Di-tyrosine residues within ITAMs are phosphorylated upon TCR engagement; their function is to recruit signaling molecules, such as protein tyrosine kinases, to the TCR complex, thereby initiating the T cell–activation cascade. Though conserved, ITAM sequences are nonidentical, raising the possibility that the diverse developmental and functional responses controlled by the TCR may be partly regulated by distinct ITAMs. We previously generated CD3-zeta–deficient and CD3-epsilon–deficient mice by gene targeting. We genetically reconstituted the mice with transgenes encoding wild-type or signaling-deficient (ITAM–mutant) forms of CD3-zeta and CD3-epsilon and characterized the developmental and functional consequences of the alterations for TCR signaling. We found that TCR–ITAMs are functionally equivalent but act in concert to amplify TCR signals and that TCR signal amplification is critical for thymocyte selection, the process by which potentially useful immature T cells are instructed to survive and differentiate further (positive selection) and by which potentially auto-reactive cells, which may cause auto-immune disease, are deleted in the thymus (negative selection).
Unexpectedly, we found that a complete complement of TCR–ITAMs is not required for most mature T cell effector functions. However, recent work showed a requirement for ITAM multiplicity for the generation of T follicular helper cells, which are required for optimal B cell antibody responses. One possible explanation for the relatively mild phenotype observed in the TCR ITAM–reduced mice is that ITAM–mediated signal amplification is not required for most mature T cell activation responses. Another is that, in ITAM–mutant mice, T cells exhibit normal functional responsiveness because of compensatory mechanisms imposed during development. To resolve this question, we recently generated a TCR–zeta chain conditional knockin mouse in which T cell development and selection can occur without attenuation of TCR signaling (i.e., in the presence of a wild-type 3-ITAM “6Y” zeta chain), but in which mature, post-selection T cells may be induced to express TCRs containing signaling-defective (0-ITAM “6F”) zeta chains in lieu of wild-type zeta chains (Figure 1). Thus, mature T cell signaling should not be influenced by potential compensatory mechanisms that operate during T cell maturation, and T cells in these mice should be faithful indicators of the role of multiple TCR ITAMs in mediating specific, mature T cell responses. Experiments with the mice confirmed that the knockin zeta locus functions as predicted. In the past year we conducted an extensive evaluation of T cell effector responses in 6F/6F and control, 6Y/6Y mice. Consistent with our previous data, we found that ‘general’ T cell responses such as cytokine production and proliferation were not significantly impaired in 6F/6F mice. However, the TCR repertoire was skewed in 6F/6F mice, as assessed by TCRValpha usage. (We stained for the spectrum of TCR alpha chain usage on mature T cells with different Valpha–specific antibodies and found that the relative percentage of each Valpha was different in the 6Y/6Y and 6F/6F mice.) Despite this, we detected no obvious ‘gaps’ in the antigen-reactive TCR repertoire in 6F/6F mice (i.e., 6F/6F mice were able to mount T cell responses to a variety of antigens and pathogens and contained similar numbers of naïve antigen-specific T cells as assessed by tetramer enrichment). The generation of memory T cell subsets was also unimpaired in 6F/6F mice. However, 6F/6F mice exhibited a significant impairment in the generation of follicular helper T cells (TFH). The fact that the same defect was observed in 6F/6F:B6CD45.1 bone marrow chimeras confirmed that it was intrinsic to T cells and not secondary to the elevated numbers of suppressor T cells (Tregs) in 6F/6F mice. After we had established that 6Y/6Y and 6F/6F mice contain similar numbers of naive 2W peptide (an immunogenic peptide)–reactive CD4 T-cells by tetramer enrichment, we observed the same TFH defect in mice infected with recombinant Listeria monocytogenes (Lm) that express a 2W peptide-OVA fusion protein (Lm-OVA), finally confirming that the defect was specifically attributable to generation of TFH cells and not a paucity of antigen-specific progenitors in 6F/6F mice. We are now using this model system to evaluate the role of ITAM multiplicity and ITAM–mediated signal amplification in T cell development, immune tolerance, and mature T cell function.
Click image to enlarge.
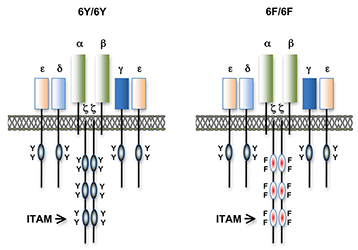
Figure 1. T cell antigen receptors expressed in 6Y/6Y and 6F/6F knock-in mice
Subunit composition of the T cell antigen receptors in 6Y/6Y and 6F/6F mice. 6Y/6Y mice express wild-type zeta chain dimers with functional ITAM signaling motifs that contain two tyrosine (Y) residues. 6F/6F mice express mutant zeta chain dimers in which the ITAM tyrosines have been changed to phenylalanine (F).
Identification and characterization of proteins important for TCR fine tuning and TCR signaling
We extended our analysis of TCR–signaling subunits to other molecules that participate in or influence the TCR–signaling response. The cell-surface protein CD5 negatively regulates TCR signaling and functions in thymocyte selection. Examination of CD5 expression during T cell development revealed that surface levels of CD5 are regulated by TCR signal intensity and by the affinity of the TCR for self-peptide ligands in the thymus that mediate selection. To determine whether the ability to regulate CD5 expression is important for thymocyte selection, we generated transgenic mice that constitutively express high levels of CD5 throughout development. Overexpression of CD5 significantly impaired positive selection of some thymocytes (those that would normally express low levels of CD5) but not of others (those that would normally express high levels of CD5). The findings support a role for CD5 in modulating TCR signal transduction and thereby influencing the outcome of thymocyte selection. Current studies are centered on identifying the mechanism by which CD5 inhibits TCR signaling and determining whether the protein's regulated expression during development is important for preventing autoimmunity. The ability of individual thymocytes to regulate CD5 expression represents a mechanism for ‘fine tuning’ the TCR signaling response during development so that the integrated signaling response can be adjusted to permit T-cell functional competency without causing autoimmunity. Reasoning that, in addition to CD5, other molecules participate in TCR tuning, we initiated microarray-based screening for genes differentially expressed in developing T cells under conditions of high- or low-affinity TCR interactions. We identified several genes from this screen for further study and are validating their function as tuning molecules. Given that the molecules regulate TCR signaling, they represent potential autoimmune-disease susceptibility markers and potential targets for treatment of patients with autoimmune disease, similar to current ‘checkpoint inhibitor’ therapies that are based on blocking the function of the induced inhibitory molecules PD-1 and CTLA-4.
Identification and characterization of Themis, a novel protein required for T cell development
Click image to enlarge.
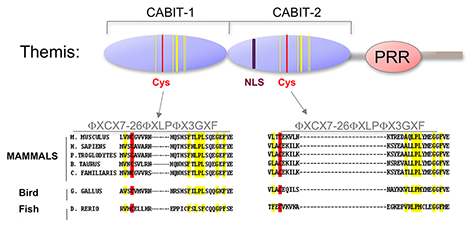
Figure 2. Themis is highly conserved in vertebrates.
Themis contains two novel CABIT domains, each with a conserved cysteine (red) and conserved flanking residues (yellow), a nuclear localization signal (NLS), and a proline-rich region (PRR).
Using a subtractive cDNA library–screening approach, we recently identified Themis, now known as Themis1, a novel T cell–specific adapter protein (Figure 2). To investigate the function of Themis1 in T cell signaling and development, we generated Themis1-knockdown cell lines: Themis1 knockout mice (conventional and conditional), and Themis1-transgenic mice. Analysis of the effects of modulating Themis1 expression revealed a critical role for the protein in late T cell development. We obtained the following results. (1) The Themis1 paralog Themis2 can substitute for Themis1 in T cell development; we found that the ability of the B cell–specific family Themis member Themis2 was equivalent to that of Themis1 to restore normal T cell development in Themis1–/– mice, thus demonstrating functional redundancy of Themis1 and Themis2. (2) We generated retroviruses encoding domain-deletion mutants of Themis1, infected Themis1–/– bone marrow progenitors, and made bone marrow chimeras to determine which regions of Themis1 are important for in vivo function; we found that the Themis1 proline-rich sequence (PRS), which mediates binding to the signaling protein Grb2, was required for in vivo function, as assessed by rescue of the developmental block in Themis1–/– thymocytes, but that the CABIT–domain cysteines are not essential. (3) We generated Themis2–/– mice and began a collaboration with Richard Cornall to characterize the mice. Our results identified an important role for Themis2 in facilitating B cell activation by low avidity, but not high avidity, B cell receptor (BCR)–antigen interactions. Themis2 was required to elicit normal Ca2+ and signaling via the Erk pathway in response to low-avidity interactions and was necessary for positive selection of B1 cells and germinal center B cells by self and foreign antigens. We detected Themis2 in complexes with the signaling proteins Grb2, Lyn, and PLCγ2 and found that it was required for normal tyrosine phosphorylation of Lyn and PLCγ2. This subtle but clear phenotype of Themis2–/– mice was not detected in a previous and less extensive study of Themis2–/–, which concluded that loss of Themis2 has no effect on B cell development or function.
Our findings show that the impact of loss of Themis1 and Themis2 on T and B cell development, respectively, is strikingly similar. In each case, the main effect is on positive selection, which is controlled by low-avidity antigen-receptor interactions. This, together with the ability of Themis2 to rescue T cell development in Themis1–/– mice, indicates that Themis1 and Themis2 perform similar functions in T and B cells. Our ongoing studies are focusing on elucidating the mechanism through which Themis participates in T cell signaling and regulates T cell development.
Role of Ldb1 transcription complexes in hematopoiesis and in T cell acute lymphoblastic leukemia
Lim domain binding protein-1 (Ldb1) is a ubiquitously expressed nuclear protein that contains a LIM–zinc finger protein–interaction motif and a dimerization domain. In hematopoietic cells, Ldb1 functions by interacting with and/or recruiting specific partners (including the LIM–only protein Lmo2 and the transcription factors SCL/Tal1 and Gata1 or Gata2) to form multi-molecular transcription complexes (Figure 3). Within the hematopoietic lineage, expression of Ldb1 is highest in progenitor cells, which include hematopoietic stem cells (HSCs). Ldb1–null (Ldb1–/–) mice die between day 9 and 10 of gestation, preventing us from directly studying the impact of loss of Ldb1 on fetal or adult hematopoiesis. We investigated the role of Ldb1 in hematopoiesis by following the fate of Ldb1–/– embryonic stem cells (ESCs) in mouse blastocyst chimeras and by conditional, stage-specific deletion of Ldb1. Significantly, Ldb1–/– ESCs were capable of generating HSCs, which could give rise to both myeloid and lymphoid lineage cells; however, the number of Ldb1–/– HSCs gradually diminished at later stages of development. Following adoptive transfer of fetal liver hematopoietic progenitor cells, Ldb1–/– HSCs were rapidly lost, indicating a failure of self-renewal or survival. More recent data indicate that the loss of Ldb1–/– HSCs results from differentiation rather than cell death. Although expressed in ESCs, Ldb1 expression is not required for ESC maintenance, indicating a selective requirement in adult stem cell populations. We performed a genome-wide screen for Ldb1–binding sites using ChIP-seq. Analysis of the ChIP-Seq data revealed that Ldb1 complexes bind at the promoter or regulatory sequences near a large number of genes known to be required for HSC maintenance. The data suggest that Ldb1 complexes function in a manner similar to Oct4/nanog/Sox2, transcription factors all essential to maintaining the pluripotent ESC phenotype, in ES cells to regulate a core transcriptional network required for adult stem cell maintenance. Examination of the function of Ldb1 in lineages downstream of the HSC identified an essential function in the erythroid lineage but not in other myeloid cells or lymphoid cells. Interestingly, ChIP-Seq analysis of Ldb1 DNA–binding complexes demonstrated that, in HSCs, Ldb1 complexes contain the transcription factor Gata2 whereas, in erythroid progenitors, Ldb1 complexes contain Gata1 (which is highly expressed in the erythroid lineage). The results indicate that multimeric Ldb1 transcription complexes have distinct functions in the hematopoietic system depending on their subunit composition, with Gata2–containing complexes regulating expression of HSC–maintenance genes and Gata1 complexes regulating expression of erythroid-specific genes (Figure 3). Current studies are aimed at investigating how Ldb1 complexes regulate gene expression and the role of Ldb1 dimerization in mediating long-range promoter-enhancer interactions in hematopoietic cells. In addition, the lab is investigating a potential role for Ldb1 in regulating self-renewal of T cell progenitors in the thymus.
Click image to enlarge.
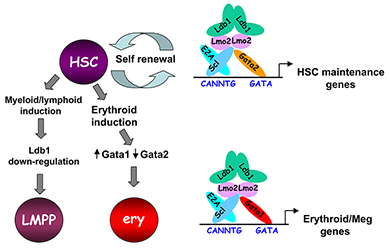
Figure 3. Model of Ldb1 function in the hematopoietic lineage
Ldb1 forms a multimeric DNA–binding complex in hematopoietic cells with the adapter Lmo2 and the transcription factors Scl and Gata1 or Gata2. In hematopoietic stem cells (HSCs), in which Gata2 is highly expressed, Ldb1-Lmo2-Scl-Gata2 complexes positively regulate expression of HSC maintenance genes. Differentiation of HSCs to the myeloid or lymphoid lineage (LMPP) is triggered by downregulation of Ldb1, whereas commitment to the erythroid lineage (ery) is triggered by induction of Gata1 and downregulation of Gata2, resulting in the formation of an Ldb1-Lmo2-Scl-Gata1 complex, which positively regulates expression of erythroid-specific genes.
Acute lymphoblastic leukemias are the most common type of cancer in children. T cell acute lymphoblastic leukemia (T-ALL) results from oncogenic transformation of immature T cell progenitors (thymocytes). Mouse models of T-ALL have been generated, and one of the most informative is the Lmo2-transgenic (Lmo2-tg) mouse, which expresses high levels of the nuclear adapter Lmo2 in thymocytes. The model closely mimics a prevalent type of human T-ALL, which is associated with chromosomal mutations that result in increased expression of LMO2. We recently reported that overexpression of Lmo2 in mouse thymocytes induces T-ALL at two distinct stages of development (an early ‘ETP’ stage and a later ‘DN3’ stage). Notably, human T-ALLs can also occur at two similar stages of thymocyte maturation. The most immature forms of T-ALL in Lmo2-tg mice and in humans express high levels of the transcription factor Hhex and are designated Early T Progenitor (ETP) T-ALL, whereas later-stage tumors are low in Hhex but express high levels of more mature markers of T cell development, including Notch1, Dtx1, Ptcra, and Hes1. Lmo2 functions as a subunit of multimeric Ldb1–nucleated DNA binding complexes. We found that normal ETP thymocyte progenitor cells express the same Ldb1 complex subunits that are present in HSCs and that ETPs exhibit HSC characteristics, including self-renewal potential. ETPs in Lmo2-tg mice appear to be ‘locked’ into a pattern of perpetual self-renewal and are refractory to normal inductive signals that promote further differentiation. Hhex is a target of Ldb1 complexes in HSCs and ETPs, a result that strongly suggests that Ldb1 complexes are responsible for the aberrant self-renewal in Lmo2-tg mice that predisposes to oncogenesis. We hypothesize that Ldb1 complexes regulate self-renewal in ETPs as well as in HSCs. Lmo2 is normally down-regulated when thymocytes undergo T-lineage commitment, suggesting that extinguishing expression of Lmo2 (and by extension, Ldb1 complexes) is important for T cell differentiation and that failure to do so predisposes to oncogenesis via ‘second-hit’ transforming events.
In preliminary RNA-Seq gene expression experiments, we found that the RNA expression signatures of Lmo2-tg immature thymocytes and HSCs are very similar, consistent with the notion that Lmo2 overexpression ‘freezes’ cells in a stem cell self-renewal state. To determine whether Ldb1 complexes are in fact required for ETP self-renewal and to explore the genes regulated by these complexes, we will determine whether Ldb1 is required for Lmo2-tg–induced thymocyte self-renewal. Such experiments will allow us to address several key questions, including whether, as predicted, Ldb1, and by extension Ldb1 complexes, regulate expression of genes that control a self-renewal genetic program in ETPs and whether Ldb1 complexes are necessary for the transcriptional/developmental effects of Lmo2. Importantly, we will also analyze the mice for T-ALL. T-ALL formation is highly penetrant in Lmo2-tg mice (virtually 100% of mice develop T-ALL by one year). We will investigate whether the timing or frequency of T-ALL is different if Ldb1 is deleted. We anticipate that these results will provide insights into the mechanisms controlling T-ALL oncogenesis in humans and may provide new therapeutic avenues for treatment of this devastating pediatric disease.
Additional Funding
- Dr. Amy Palin is applying for an NIH K99 award.
- NICHD Director's Award
Publications
- Choi YS, Gullicksrud JA, Xing S, Zeng Z, Shan Q, Li F, Love PE, Peng W, Xue HH, Crotty S. LEF-1 and TCF-1 orchestrate TFH differentiation by regulating differentiation circuits upstream of the transcriptional repressor Bcl6. Nat Immunol 2015;16:980-990.
- Hwang S, Palin AC, Li L, Song KD, Lee J, Herz J, Tubo N, Chu H, Pepper M, Lesourne R, Zvezdova E, Pinkhasov J, Jenkins MK, McGavern D, Love PE. TCR ITAM multiplicity is required for the generation of follicular helper T-cells. Nat Commun 2015;6:6982.
- Cheng D, Deobagkar-Lele M, Zvezdova E, Choi S, Lesourne R, Uehara S, Baup D, Bennett SC, Bull KR, Crockford TL, Ferry H, Anzilotti C, Love PE, Cornall RJ. Themis2 lowers the threshold for B cell activation during positive selection. Nat Immunol 2017;18(2):205-213.
- Kitmura MY, Thomas J, Tai X, Guinter TI, Shinzawa M, Etzensperger R, Li Z, Love P, Nalayama T, Singer A. Timing and duration of MHC I positive selection signals are adjusted in the thymus to prevent lineage errors. Nat Immunol 2016;17(12):1415-1423.
- Zvezdova E, Mikolajczak J, Garreau A, Marcellin M, Rigal L, Lee J, Choi S, Blaize G, Argenty J, Familiades J, Li L, Gonzalez de Peredo A, Burlet-Schiltz O, Love PE, Lesourne R. Themis1 enhances T cell receptor signaling during thymocyte development by promoting Vav1 activity and Grb2 stability. Sci Signal 2016;9(428):ra51.
- Triplett TA, Cardenas KT, Lancaster JN, Hu Z, Selden HJ, Jasso GJ, Balasubramanyam S, Chan K, Li L, Chen X, Marcogliese AN, Dave UP, Love PE, Ehrlich LI. Endogenous dendritic cells from the tumor microenvironment support T-ALL growth via IGF1R activation. Proc Natl Acad Sci USA 2016;113(8):E1016-1025.
Collaborators
- Remy Bosselut, PhD, Laboratory of Immune Cell Biology, NCI, Bethesda, MD
- Richard J. Cornall, FMedSci, FRCP, University of Oxford, Oxford, United Kingdom
- Utpal P. Davé, MD, Vanderbilt University Medical Center, Nashville, TN
- Lauren Ehrlich, PhD, The University of Texas, Austin, TX
- Marc Jenkins, PhD, University of Minnesota, Minneapolis, MN
- Dorian McGavern, PhD, Viral Immunology and Intravital Imaging Section, NINDS, Bethesda, MD
- Alfred Singer, MD, Experimental Immunology Branch, NCI, Bethesda, MD
- Nan-ping Weng, PhD, Laboratory of Immunology, NIA, Baltimore, MD
Contact
For more information, email lovep@mail.nih.gov or visit irp.nih.gov/pi/paul-love.


