Regulation of Mammalian Cell Proliferation and Differentiation

- Melvin L. DePamphilis, PhD, Head, Section on Eukaryotic Gene Regulation
- Kotaro Kaneko, PhD, Staff Scientist
- Alex Vassilev, PhD, Staff Scientist
- Diane Adler-Wailes, MS, Senior Research Assistant
- Xiaohong Zhang, BA, Technical Assistant
- Christelle de Renty, PhD, Visiting Fellow
- Lori Griner, PhD, Visiting Fellow
- Yi-Yuan Huang, PhD, Visiting Fellow
- Courtney Kurtyka, PhD, Visiting Fellow
Nothing is more fundamental to living organisms than the ability to reproduce. Each time a human cell divides, it must duplicate its genome, a problem of biblical proportions. A single fertilized human egg contains 2.1 meters of DNA. An adult of about 75 kg (165 lb) consists of about 29 trillion cells containing a total of about 60 trillion meters of DNA, a distance equal to 400 times the distance from Earth to sun. Not only must the genome be duplicated trillions of times during human development, but must be duplicated once and only once each time a cell divides (termed mitotic cell cycles). If we interfere with this process by artificially inducing cells to re-replicate their nuclear genome before cell division, the result is DNA damage, mitotic catastrophe, and programmed cell death (apoptosis). On rare occasions, specialized cells can duplicate their genome several times without undergoing cell division (termed endocycles), but when this occurs, it generally results in terminally differentiated polyploid cells that are viable but no longer proliferate. As we age, however, the ability to regulate genome duplication diminishes, resulting in genome instability, which allows genetic alterations that can result in promiscuous cell division—better known as cancer. For a comprehensive description of genome duplication in all forms of life, refer to Genome Duplication (Figure 2).
Our research program focuses on three questions. What are the mechanisms that restrict genome duplication to once per cell division? How are these mechanisms circumvented to allow developmentally programmed induction of polyploidy in terminally differentiated cells? How can we manipulate these mechanisms to destroy cancer cells selectively?

Figure 1. DNA Replication (Cold Spring Harbor Perspectives in Biology, 2013), edited by Marcel Mechali, Melvin L. DePamphilis, and Stephen D. Bell
The collection covers all aspects of DNA replication and its control across all domains of life. The contributors examine the molecular machinery involved in the assembly of replication origin complexes, the establishment of replication forks, unzipping of the double helix, priming of DNA synthesis, and elongation of daughter strands. Chromatin organization and dynamics, lagging-strand maturation, telomere replication, and mechanisms to handle errors and damage in DNA are also discussed.

Click image to enlarge.
Figure 2. Genome Duplication (Garland Science, 2010) by Melvin L. DePamphilis and Stephen D. Bell
The book contains the first comprehensive description of the subject since Kornberg and Baker’s book DNA Replication, second edition (W.H. Freeman, 1992). Genome Duplication describes the principles, concepts, and mechanisms common to genome duplication in all three domains of life: bacteria, archaea, and eukarya. In addition, it discusses the evolution of the DNA replication machinery and the impact of DNA replication on human disease.
Regulation of DNA replication in mammalian cells
Genome duplication begins when the six-subunit ORC (origin recognition complex) binds to specific chromosomal loci termed origins of bidirectional replication, which we and others mapped at specific sites in the genomes of flies and mammals. The sites are determined by both genetic and epigenetic features. The number and location of replication origins in the cells of multicellular organisms can change from an average of one in every 10 to 20 kb in the rapidly cleaving embryos of frogs, flies, and fish to one in every 50 to 300 kb in the differentiated cells of adult organisms. Developmental changes in origin density also occur during specific stages in animal development. Thus, metazoan genomes contain many potential replication origins but, during development, some of these sites are selectively activated while others are suppressed, a concept introduced many years ago as the "Jesuit Model", because many are called, but few are chosen (DePamphilis ML, Ann Rev Biochem 1993;62:29; Curr Opin Cell Biol 1993;5:434; J Biol Chem 1993;268:1).
ORC initiates assembly of prereplication complexes (preRCs) consisting of a DNA helicase loader [ORC(1–6) and replication factors Cdc6 and Cdt1] and the replicative DNA helicase [Mcm(2–7)]. Several years ago, we discovered that the behavior of ORC in mammalian cells differs significantly from that in single-cell eukaryotes such as yeast. In contrast to yeast, Orc1 associates weakly with the stable core complex ORC(2–5), and the ability of ORC to initiate DNA replication depends on this interaction. Moreover, it appears that the interaction of Orc1 with ORC(2–5) is one of the mechanisms that regulate when and where initiation events occur. We termed this concept the ORC cycle, and we and others have established its basic features (Figure 3). Cell cycle–dependent modifications of Orc1 regulate Orc1 activity, and Orc1 activity regulates ORC activity, which regulates initiation of DNA replication.
Click image to enlarge.
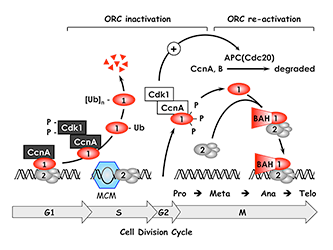
Figure 3. The ORC cycle in mammalian cells (see Noguchi et al., EMBO J 2006;25:5372 and references therein)
ORC(1–6) is bound to chromatin during the G1 phase of the cell cycle, at which time it is part of a prereplication complex. When S phase begins, the association between Orc1 and chromatin-bound ORC(2–6) is destabilized by selective CDK–dependent phosphorylation and ubiquitination. Monoubiquitinated Orc1 is exported to the cytoplasm. The remaining ORC subunits are subsequently released from chromatin. Polyubiquitinated Orc1 is degraded by the 26S proteasome. Orc1 levels are restored during the G2-to-M transition, but Orc1 is hyperphosphorylated, an event that prevents ORC assembly. During the anaphase-to-G1 phase transition, Orc1 is dephosphorylated and, together with other ORC subunits, binds to chromatin, an event facilitated by the Orc1 BAH domain. If Orc1 is not associated with other ORC subunits, or if it is not phosphorylated or ubiquitinated, it induces apoptosis.
The ORC cycle is only one of six known mechanisms that can determine when and where DNA replication begins in human cells. Cdk2•CcnA (cyclin-dependent kinase-2•cyclin A) also suppresses Cdc6 (cell vision cycle-6) and Cdt1 (chromatin licensing and DNA replication factor-1) activities by phosphorylation, and Cdt1 is targeted by two ubiquitin ligases and by geminin, a Cdt1–specific protein inhibitor (Figure 4). However, not every pathway is active in all cell types. For example, cells derived from human cancers are dependent on geminin to prevent DNA re-replication, whereas cells derived from normal human tissues are dependent on both geminin and cyclin A–dependent CDK (cyclin-dependent kinase) activity.
Click image to enlarge.
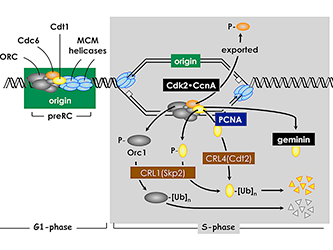
Figure 4. Several convergent pathways restrict genome duplication to once per cell division.
In human cells, activity of the replication factor Cdt1 is down-regulated in four ways. Free Cdt1 is phosphorylated by Cdk2 in association with Cyclin A (CcnA), thereby suppressing Cdt1 activity and converting Cdt1 into a substrate for the CRL1(Skp2) ubiquitin ligase. As replication forks pass through the origin, Cdt1 binds to the proliferating cell nuclear antigen (PCNA) clamp, which holds the replicative DNA polymerase onto the replication fork. The Cdt1•PCNA•chromatin form of Cdt1 is a substrate for the CRL4(Cdt2) ubiquitin ligase. Ubiquitinated Cdt1 is then degraded by the 26S proteasome. Finally, geminin binds to Cdt1 and inhibits its activity. In addition to inactivation of Cdt1, CcnA–dependent phosphorylation of Orc1 prevents it from binding to chromatin during mitosis, and CcnA–dependent phosphorylation of Cdc6 suppresses its activity and promotes its nuclear export.
A rare event in mammals, endoreduplication, by which the nuclear portion of the genome is duplicated one or more times (endocycles) without an intervening mitosis, is a common event among arthropods and plants. In mammals, it first occurs during peri-implantation development when cells within the trophectoderm (TE) of the blastocyst are deprived of the mitogenic factor FGF4 (fibroblast growth factor-4). Trophoblast stem (TS) cells then differentiate into the polyploid, viable, nonproliferating trophoblast giant (TG) cells required for embryo implantation and placentation (Figure 5). PreRC assembly requires the absence of both CDK activity and geminin, a condition that occurs in mitotic cell cycles during the anaphase-to-G1 phase transition. We discovered that this condition could be induced in TS cells by selective chemical inhibition of CDK1, the enzyme required for entry into mitosis, but not in embryonic stem (ES) cells, in which the consequence is apoptosis. Therefore, selective inhibition of CDK1 triggers endoreduplication only in cells programmed to differentiate into polyploid cells. Similarly, FGF4 deprivation of TS cells induced expression of two CDK–specific inhibitors: p57/Kip2 and p21/Cip1. One (p57) was essential for endoreduplication while the other (p21) appeared to facilitate p57 activity by suppressing expression of Chk1 (checkpoint kinase-1) and the mitotic inhibitor Emi1 (Figure 6). TS cells (+FGF4) express both the p57 and p21 genes, but the nonactivated form of Chk1 phosphorylates the p57 and p21 proteins, thereby targeting them for ubiquitin-dependent degradation. In TG cells, CHK1 is suppressed to allow p57 expression during G phase, CDK2 is required for DNA replication, and p57 is degraded during S phase to allow endocycles.
Click image to enlarge.
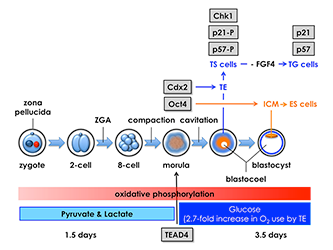
Figure 5. Mouse pre-implantation development: zygotic gene activation (ZGA) begins at the two-cell stage.
The eight totipotent blastomeres at the eight-cell stage compact into a morula, and transcription factors Tead4, Cdx2, and Oct4 mark the beginning of a chain of events that specifies the trophectoderm (TE) and inner cell mass (ICM). Oct4 is the first gene whose expression is essential for maintaining blastomeres in a totipotent state, thereby producing the ICM and embryonic stem (ES) cells that differentiate into embryonic tissues. Tead4 is the first gene whose expression is essential for differentiation of blastomeres into TE and trophoblast stem (TS) cells, which produce the placenta. In the absence of Tead4 expression, all totipotent blastomeres produce OCT4 protein, revealing that the TEAD4 protein is required for Cdx2 gene expression. However, TEAD4 is essential for blastocyst formation only under conditions that increase the rate of oxidative phosphorylation. Following compaction, pre-implantation embryos switch energy substrates from pyruvate and lactate to glucose in order to meet the increased energy demands of blastocoel formation, an event unique to the TE. Thus, Tead4 does not specify the TE; rather, TEAD4 maintains energy homeostasis so that other genes can specify the TE.
Click image to enlarge.
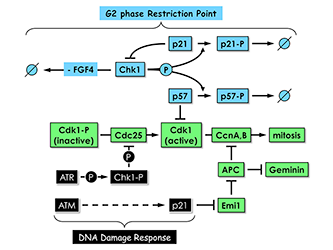
Figure 6. Proliferating trophoblast stem (TS) cells exhibit a G2 phase restriction point analogous to the G1 restriction point in proliferating cultured mammalian cells.
In the presence of the mitogen FGF4 and in the absence of DNA damage, Chk1, the same checkpoint kinase that is activated by DNA damage, phosphorylates both p57 and p21, thereby targeting them for ubiquitin-dependent degradation. In the absence of FGF4 and DNA damage, Chk1 expression is suppressed by an as yet unidentified mechanism. Both p57 and p21 proteins accumulate; p57 inhibits CDK1, and TS cells exit their mitotic cell cycle and differentiate into trophoblast giant (TG) cells; and p21 plays several roles, including suppressing Chk1 expression to maintain TG cell status and inhibiting Emi1 to activate the anaphase-promoting complex (APC), which promotes preRC assembly by targeting cyclins A and B and geminin for degradation. Both Chk1 and p21 are also components of the DNA–damage response that prevents mitosis until the problem is corrected; the features that distinguish their roles in the two regulatory mechanisms remain to be elucidated.
DNA replication and cell differentiation during pre-implantation development
Fertilization activates the first round of genome duplication, after which the fertilized egg cleaves into a two-cell embryo (Figure 5). In mice, the two-cell embryo then activates expression of about 300 genes that are required to continue development of the organism (termed zygotic gene activation). Initially, every cell (blastomere) produced by the cleavage events is 'totipotent,' that is, it is capable of giving rise to the entire organism. But, within five rounds of cell cleavage, a blastocyst appears, marking the beginning of cell differentiation. The blastocyst consists of a spherical monolayer of epithelial cells called the TE that gives rise to TS cells, TG cells, and eventually to the placenta. The TE layer encompasses a group of cells called the inner cell mass that gives rise to ES cells and eventually to the embryo.
Our goal for some years has been to determine whether the requirements for genome duplication in cultured cells were the same for cleavage-stage embryos, for the stage prior to cell differentiation, and for the transition from mitotic cell cycles to endocycles. To this end, we identified genes that we thought might be critical for pre-implantation development. The results have been fruitful, but surprising. We discovered that Dkkl1, a gene unique to mammals, is expressed specifically during implantation of the embryo and development of spermatocytes into sperm. Moreover, we showed that inactivation of Dkkl1 in mice resulted in the production of sperm that are defective in fertilization. We discovered that Tead2, one of a highly conserved family of four transcription factors that share a common DNA–binding domain, is expressed from the two-cell embryo throughout pre-implantation development but, remarkably, is not required until after implantation and the start of nervous system formation. Mice lacking a functional Tead2 gene have difficulty forming a neural tube. Failure to close the neural tube in mice is called exencephaly, which is related to anencephaly, the common human birth defect that can be prevented by folic acid. Finally, we discovered that Tead2 and Tead4 are the only Tead genes expressed in pre-implantation mouse embryos and that, in contrast to genetic inactivation of Tead2, genetic inactivation of Tead4 results in the formation of a morphologically abnormal morula, the absence of TE–specific genes, and failure to produce a blastocoel. These and other results in the literature strongly suggested that Tead4 is a master gene that sets in motion the first round of cell differentiation during mammalian development. Surprisingly, our subsequent research demonstrated otherwise.
Master genes trigger a sequence of events that specify cell fate. However, genes essential for maintaining a metabolic state that allows other genes to specify cell fate may be mistaken for master genes. We validated the concept by showing that Tead4, the presumptive master gene for trophectoderm specification, is a homeostatic gene that is essential for blastocoel formation in utero, a prerequisite for embryo implantation and placentation. Requirement for Tead4 is induced metabolically, not developmentally, and conditions that minimize oxidative stress allow embryos lacking TEAD4 to develop into blastocysts expressing genes required to produce functional trophectoderm. TEAD4, which is expressed at the eight-cell stage, is essential only under conditions that promote energy production by oxidative phosphorylation (Figure 5). Given that either inhibition of the mTOR pathway or an anti-oxidant alleviates the requirement, TEAD4 appears to regulate changes in redox potential that are expected to occur during the high levels of oxidative phosphorylation associated with blastocyst formation in utero (Kaneko and DePamphilis, Development 2013;140:3680-3689).
Geminin prevents trophoblast stem cells from exiting mitosis and differentiating into the nonproliferating, viable, giant cells essential for implantation and placentation.
The protein geminin is involved in both DNA replication and cell fate acquisition. Although it is essential for mammalian preimplantation development, its role remains unclear. In one study, ablation of the geminin gene (Gmnn) in mouse preimplantation embryos resulted in apoptosis, suggesting that geminin prevents DNA re-replication, whereas in another study it resulted in differentiation of blastomeres into TG cells, suggesting that geminin regulates trophoblast specification and differentiation. Other studies led to the conclusion that trophoblast differentiation into TG cells is regulated by FGF4 and that geminin is required to maintain endocycles. We showed that ablation of Gmnn in TS cells proliferating in the presence of FGF4 closely mimics the events triggered by FGF4 deprivation: arrest of cell proliferation, formation of giant cells, excessive DNA replication in the absence of DNA damage and apoptosis, and changes in gene expression that include loss of the checkpoint kinase Chk1 with up-regulation of p57 and p21. Moreover, FGF4 deprivation of TS cells reduces geminin to a basal level that is required for maintaining endocycles in TG cells. Thus, geminin acts like a component of the FGF4 signal transduction pathway that governs trophoblast proliferation and differentiation, and the protein is required to maintain endocycles (de Renty, Kaneko and DePamphilis, Dev Biol 2014;387:49-63).
Geminin is essential to prevent DNA re-replication–dependent apoptosis in pluripotent cells, but not in differentiated cells.
Geminin is a dual-function protein unique to multicellular animals with roles in modulating gene expression and preventing DNA re-replication. We showed that geminin is essential at the beginning of mammalian development to prevent DNA re-replication in pluripotent cells, exemplified by ES cells, as they undergo self-renewal and differentiation. ES cells, embryonic fibroblasts, and immortalized fibroblasts were characterized before and after geminin was depleted either by gene ablation or siRNA. Depletion of geminin under conditions that promote either self-renewal or differentiation rapidly induced DNA re-replication, followed by DNA damage, then a DNA–damage response, and finally apoptosis. Once differentiation had occurred, geminin was no longer essential for viability, although it continued to contribute to prevent DNA re-replication–induced DNA damage. We detected no relationship between expression of geminin and genes associated with either pluripotency or differentiation. Thus, the primary role of geminin at the beginning of mammalian development is to prevent DNA re-replication–dependent apoptosis, a role previously believed essential only in cancer cells. The results suggest that regulation of gene expression by geminin occurs only after pluripotent cells differentiate into cells in which geminin is not essential for viability (Reference 1).
Cytoplasmic localization of p21 protects trophoblast giant cells from DNA damage–induced apoptosis.
Proliferating TS cells can differentiate into nonproliferating, but viable, TG cells that are resistant to DNA damage–induced apoptosis. Differentiation is associated with selective up-regulation of the Cip/Kip cyclin–dependent kinase inhibitors p57 and p21, while expression of p27 remains constant. Previous studies showed that p57 localizes to the nucleus in TG cells, where it is essential for endoreplication. We showed that p27 also remains localized to the nucleus during TS cell differentiation, where it complements the role of p57. Unexpectedly, p21 localized to the cytoplasm, where it was maintained throughout both the G- and S-phases of endocycles, and where it prevented DNA damage–induced apoptosis. This unusual status for a Cip/Kip protein was dependent on site-specific phosphorylation of p21 by the Akt1 kinase that is also up-regulated in TG cells. Although cytoplasmic p21 is widespread among cancer cells, among normal cells it has been observed only in monocytes. The fact that it also occurs in TG cells reveals that p57 and p21 serve nonredundant functions and suggests that the role of p21 in suppressing apoptosis is restricted to terminally differentiated cells (Reference 2).
And-1 coordinates with Claspin for efficient Chk1 activation in response to replication stress.
The replisome is important for DNA replication checkpoint activation, but how specific components of the replisome coordinate with the ATR kinase to activate Chk1 in human cells remains largely unknown. We demonstrated that And-1, a replisome component, acts together with ATR to activate Chk1. And-1 is phosphorylated at T826 by ATR following replication stress, and the phosphorylation is required for And-1 to accumulate at the damage sites, where And-1 promotes the interaction between Claspin and Chk1, thereby stimulating efficient Chk1 activation by ATR. Significantly, And-1 binds directly to ssDNA (single-strand DNA) and facilitates the association of Claspin with ssDNA. Furthermore, And-1 associates with replication forks and is required for the recovery of stalled forks. The studies establish a novel ATR–And-1 axis as an important regulator for efficient Chk1 activation and reveal a novel mechanism by which the replisome regulates the replication checkpoint and genomic stability (Reference 3).
Publications
- Huang YY, Kaneko KJ, Pan H, DePamphilis ML. Geminin is essential to prevent DNA re-replication-dependent apoptosis in pluripotent cells, but not in differentiated cells. Stem Cells 2015; 33(11):3239-3253.
- de Renty C, DePamphilis ML, Ullah Z. Cytoplasmic localization of p21 protects trophoblast giant cells from DNA damage induced apoptosis. PLoS One 2014; 9:e97434.
- Hao J, de Renty C, Li Y, Xiao H, Kemp MG, Han Z, DePamphilis ML, Zhu W. And-1 coordinates with Claspin for efficient Chk1 activation in response to replication stress. EMBO Journal 2015; 34:2096-2110.
- Kohn MJ, DePamphilis ML. Derivation of mouse trophoblast stem cells from preimplantation embryos. NIH Intramural Pluripotent Stem Cell Protocol Book 2014; 1:1-20.
- Ullah Z, de Renty C, DePamphilis ML. Propagation and differentiation of mouse trophoblast stem cells. NIH Intramural Pluripotent Stem Cell Protocol Book 2014; 1:21-41.
Collaborators
- Scott E. Martin, PhD, NIH Chemical Genomics Center, NHGRI, Rockville, MD
- Wenge Zhu, PhD, George Washington University Medical School, Washington, DC
Contact
For more information, email depamphm@mail.nih.gov or visit http://depamphilislab.nichd.nih.gov.


