Regulatory Small RNAs and Small Proteins

- Gisela Storz, PhD, Head, Section on Environmental Gene Regulation
- Aixia Zhang, PhD, Staff Scientist
- Tamira K. Butler-Lively, PhD, Postdoctoral Fellow
- Michael D. Dambach, PhD, Postdoctoral Fellow
- Yue Hao, PhD, Postdoctoral Fellow
- Andrew B. Kouse, PhD, Postdoctoral Fellow
- Medha V. Raina, PhD, Postdoctoral Fellow
- Taylor B. Updegrove, PhD, Postdoctoral Fellow
- Jeremy S. Weaver, PhD, Postdoctoral Fellow
- Aracely A. Romero, BS, Postbaccalaureate Fellow
- Hanbo Wang, BS, Graduate Student
Currently, we have two main interests: identification and characterization of small noncoding RNAs and identification and characterization of small proteins of less than 50 amino acids. Both small RNAs and small proteins have been overlooked because they are not detected in biochemical assays and the corresponding genes are poorly annotated and missed in genetic screens. However, mounting evidence suggests that both classes of small molecules play important regulatory roles.
Identification and characterization of small regulatory RNAs
During the past 15 years, we have carried out several different systematic screens for small regulatory RNA genes in Escherichia coli. The screens have included computational searches for conservation of intergenic regions and direct detection after size selection or co-immunoprecipitation with the RNA–binding protein Hfq. We recently examined small RNA expression using deep sequencing to further extend our identification of small RNAs, particularly antisense RNAs (Reference 1).
A major focus of the group has been to elucidate the functions of the small RNAs that we and others have identified. Early on, we showed that the OxyS RNA, whose expression is induced in response to oxidative stress, acts to repress translation through limited base pairing with target mRNAs. We discovered that OxyS action is dependent on the Sm-like Hfq protein, which functions as a chaperone to facilitate OxyS RNA base pairing with its target mRNAs. Recently, we carried out an extensive mutational studies of Hfq (Reference 2). The analysis revealed that amino acids on three different RNA interaction surfaces—the proximal face, the distal face, and the rim of the ring-shaped hexamer—differentially impact Hfq association with small RNAs and their mRNA targets (Figure 1).
It is now clear that Hfq–binding small RNAs, which act through limited base pairing, are integral to many different stress responses in E. coli. For example, we showed that the Spot 42 RNA, whose levels are highest when glucose is present, plays a broad role in catabolite repression by directly repressing genes involved in central and secondary metabolism, redox balancing, and the consumption of diverse non-preferred carbon sources. It was previously reported that the transcription factor Sigma(E) maintains membrane homeostasis in part by inducing synthesis of two small regulatory RNAs that down-regulate synthesis of abundant membrane porins. We discovered a third Sigma(E)-dependent small RNA, MicL, that is transcribed from a promoter located within the coding sequence of the cutC gene (Reference 3). MicL possesses features typical of Hfq–binding small RNAs but surprisingly targets only a single mRNA, which encodes the outer membrane lipoprotein Lpp, the most abundant protein of the cell. Interestingly, we found that the copper-sensitivity phenotype previously ascribed to inactivation of the cutC gene is actually derived from the loss of MicL and elevated Lpp levels. The observation raised the possibility that other phenotypes currently attributed to protein defects are the result of deficiencies in unappreciated regulatory RNAs and prompted us to ask questions about the evolution of base-pairing small RNAs (Updegrove TB, Shabalina SA, Storz G, FEMS Microbiol Rev 2015;39:379).
In addition to small RNAs that act via limited base pairing, we have been interested in regulatory RNAs that act by other mechanisms. For example, early work showed that the 6S RNA binds to and modulates RNA polymerase by mimicking the structure of an open promoter. In a more recent study, we discovered that a broadly conserved RNA structure motif, the yybP-ykoY motif, found in the 5′ UTR of the mntP gene encoding a manganese exporter, directly binds to manganese, resulting in a conformation that liberates the ribosome-binding site (Reference 4). Remarkably, we were able to recapitulate the manganese-dependent activation of translation in vitro. We also found that the yybP-ykoY motif responds to manganese ions in Bacillus subtilis. The identification of the yybP-ykoY motif as a manganese ion sensor suggests that the genes that are preceded by this motif and that encode a diverse set of poorly characterized membrane proteins have roles in metal homeostasis.
Studies to further characterize other Hfq–binding RNAs, antisense RNAs, and small RNAs that act in ways other than base pairing are ongoing.
Click image to enlarge.
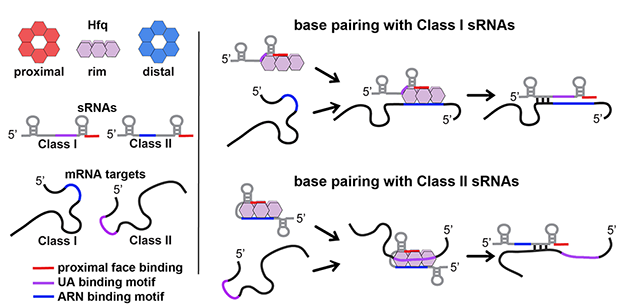
Figure 1. Model of RNA binding to Hfq
The cartoon model of the Hfq hexamer depicts the three RNA binding surfaces of Hfq: proximal face (red), rim (purple), and distal face (blue). For sRNAs and mRNAs, elements in red, purple and blue represent sequences that bind to the proximal face, the rim, and the distal face, respectively. The model depicts two alternative pathways for binding and regulation by sRNA:mRNA pairs. Class I sRNAs utilize a U–rich rho-independent terminator for binding to the proximal face and a UA binding motif for interaction with the rim. mRNA targets regulated by this class of sRNAs utilize ARN binding motifs for interacting with the distal face of Hfq. Class II sRNAs utilize the U–rich rho-independent terminator for binding to the proximal face of Hfq and an ARN binding motif in the 5′ end of the sRNA for binding to the distal face. mRNA targets regulated by this class of sRNAs contain UA–binding motifs that allow for binding to the rim of Hfq.
Identification and characterization of small proteins
In our genome-wide screens for small RNAs, we found that several short RNAs encode small proteins. The correct annotation of the smallest proteins is one of the greatest challenges of genome annotation, and perhaps more importantly, few annotated short ORFs have been confirmed to correspond to synthesized proteins. Although these proteins have largely been missed, the few small proteins that have been studied in detail in bacterial and mammalian cells have been shown to have important functions in signaling and in cellular defenses (Storz et al., Annu Rev Biochem 2014;83:18.1-18.25). We thus established a project to identify and characterize proteins of less than 50 amino acids.
We used sequence conservation and ribosome binding site models to predict genes encoding small proteins, defined as having 16–50 amino acids, in the intergenic regions of the model E. coli genome. We tested expression of these predicted as well as previously annotated small proteins by integrating the sequential peptide affinity tag directly upstream of the stop codon on the chromosome and assaying for synthesis using immunoblot assays. The approach confirmed that 20 previously annotated and 18 newly discovered proteins of 16–50 amino acids are synthesized. We have now initiated complementary biochemical approaches to identify additional small proteins.
Remarkably, more than half the newly discovered proteins are predicted to be transmembrane proteins, an observation that prompted us to examine the localization, topology, and membrane insertion of the small proteins. Biochemical fractionation showed that, consistent with the predicted transmembrane helix, the small proteins are generally most abundant in the inner membrane fraction. We found examples of both Nin-Cout (N-terminalin–C-terminalout) and Nout-Cin orientations as well as dual topology in assays of topology-reporter fusions to representative small transmembrane proteins. In addition, fractionation analysis of small protein localization in strains depleted of SecE or YidC uncovered differential requirements for these membrane insertion pathways. Thus, despite their diminutive size, small proteins display considerable diversity in topology, biochemical features, and insertion pathways.
We now are employing many of the approaches the group has used to characterize the functions of small regulatory RNAs to elucidate the functions of the small proteins. Systematic assays for the accumulation of tagged versions of the proteins showed that many small proteins accumulate under specific growth conditions or after exposure to stress. We also generated and screened bar-coded null mutants and identified small proteins required for resistance to cell-envelope stress and acid shock. In addition, the attached sequential peptide affinity tag is being exploited to identify co-purifying complexes. The combination of these approaches is giving insights into when, where, and how the small proteins are acting.
For example, we found that synthesis of a 42-amino acid protein, now denoted MntS (formerly the small RNA gene rybA), is repressed by high levels of manganese through MntR. In recent studies, we showed that MntS helps manganese activate a variety of enzymes under manganese-poor conditions, while overproduction of MntS leads to very high intracellular manganese and bacteriostasis under manganese-rich conditions (Reference 5). These and other phenotypes led us to propose that MntS modulates intracellular manganese levels, possibly by inhibiting the manganese exporter MntP.
We also discovered that the 49–amino acid inner membrane protein AcrZ (formerly named YbhT) associates with the AcrAB-TolC multidrug efflux pump, which confers resistance to a wide variety of antibiotics and other compounds. Co-purification of AcrZ with AcrB, in the absence of both AcrA and TolC, two-hybrid assays, and suppressor mutations indicate that the interaction occurs through the inner membrane protein AcrB. Mutants lacking AcrZ are sensitive to many of, but not all, the antibiotics transported by AcrAB-TolC. Such differential antibiotic sensitivity suggests that AcrZ may enhance the ability of the AcrAB-TolC pump to export certain classes of substrates. The work, together with our ongoing studies of other small proteins, suggests that many are acting as regulators of larger membrane protein complexes.
Publications
- Thomason MK, Bischler T, Eisenbart SK, Förstner KU, Zhang A, Herbig A, Nieselt K, Sharma CM, Storz G. Global transcriptional start site mapping using dRNA-seq reveals novel antisense RNAs in Escherichia coli. J Bacteriol 2015; 197:18-28.
- Schu DJ, Zhang A, Gottesman S, Storz G. Alternative Hfq-sRNA interaction modes dictate alternative mRNA recognition. EMBO J 2015; 34:2557-2573.
- Guo MS, Updegrove TB, Gogol EB, Shabalina SA, Gross CA, Storz G. MicL, a new sE-dependent sRNA, combats envelope stress by repressing synthesis of Lpp, the major outer membrane lipoprotein. Genes Dev 2014; 28:1620-1634.
- Dambach M, Sandoval M, Updegrove TB, Anantharaman V, Aravind L, Waters LS, Storz G. The ubiquitous yybP-ykoY riboswitch is a manganese-responsive regulatory element. Mol Cell 2015; 57:1099-1109.
- Martin JE, Waters LS, Storz G, Imlay JA. The Escherichia coli small protein MntS and exporter MntP optimize the intracellular concentration of manganese. PLoS Genet 2015; 11:e1004977.
Collaborators
- Susan Gottesman, PhD, Laboratory of Molecular Biology, NCI, Bethesda, MD
- Carol A. Gross, PhD, University of California San Francisco, San Francisco, CA
- James A. Imlay, PhD, University of Illinois, Urbana, IL
- Aravind L. Iyer, PhD, National Center for Biotechnology Information, NIH, Bethesda, MD
- Kay Nieselt, PhD, Universität Tübingen, Tübingen, Germany
- Kumaran S. Ramamurthi, PhD, Laboratory of Molecular Biology, NCI, Bethesda, MD
- Svetlana A. Shabalina, PhD, National Center for Biotechnology Information, NIH, Bethesda, MD
- Cynthia M. Sharma, PhD, Research Centre for Infectious Diseases, Universität Würzburg, Germany
- Lauren S. Waters, PhD, University of Wisconsin, Oshkosh, WI
- Yuri I. Wolf, PhD, National Center for Biotechnology Information, NIH, Bethesda, MD
Contact
For more information, email storz@helix.nih.gov or visit http://cbmp.nichd.nih.gov/segr.


