Signaling and Secretion in Neuroendocrine Cells

- Stanko S. Stojilkovic,
PhD, Head, Section on Cellular Signaling - Stephanie Constantin, PhD, Staff Scientist
- Naseratun Nessa, PhD, Visiting Fellow
- Kunal Pratap, PhD, Visiting Fellow
- Sonja Sivcev, PhD, Visiting Fellow
- Kosara Smiljanic, PhD, Visiting Fellow
The Section was formed in 1993 to investigate cascades of cell signaling, gene expression, and secretion in hypothalamic and pituitary cells, with a particular emphasis on the interactions between electrical events at the plasma membrane and receptor-controlled pathways. Specifically, we are concerned with how these neuroendocrine cells use ion channels and G protein–coupled receptors (GPCRs) as signaling platforms for efficient information processing. To this end, we characterize both native and recombinant receptors and channels from neuroendocrine cells. In the past, our work focused on the role of inositol-trisphosphate receptors in the oscillatory calcium release and influx, the mechanism of periodic activation of these channels, and the complex way in which calcium release from intracellular stores is synchronized with the electrical activity of the cells. We also characterized voltage-gated channels expressed in neuroendocrine cells, the cell type–specific electrical activity patterns and channels involved, the physiological significance of such activity, and the crosstalk between GPCRs and ion channels. More recently, we characterized ligand-gated receptor channels expressed in pituitary cells, including ATP–dependent P2X receptor channels. Our current work focuses on age-, sex-, and tissue-specific pituitary signaling, transcription, and secretion, the heterogeneity of pituitary secretory cells reflecting their postnatal genesis, and cell type–specific exocytic pathways. Ongoing and proposed projects include the use of conditional knockout mouse models. Results already obtained in these projects and those from other studies have revealed the great complexity and physiological significance of the intracellular signaling system in the pituitary and the need for further research. The information we are currently obtaining and the planned research should continue to be useful to researchers and clinicians in the neuroendocrine field, as well as the broader scientific community.
Somatostatin is a common agonist for most endocrine pituitary cell types.
Somatostatin receptors (SSTRs) belong to a receptor family consisting of five members, designated SSTR1–SSTR5, but the expression of these receptors within pituitary endocrine cells and their specific roles in the excitability and voltage-gated calcium signaling have not been fully characterized. To identify which subtypes of these receptors are expressed in pituitary endocrine cells, we used scRNA-Seq analysis of freshly dispersed rat pituitary cells. This analysis revealed expression of the receptor genes Sstr1, Sstr2, Sstr3, and Sstr5 in a cell type–specific manner: Sstr1 and Sstr2 in thyrotrophs, Sstr3 in gonadotrophs and lactotrophs, Sstr2, Sstr3, and Sstr5 in somatotrophs, and none in corticotrophs or melanotrophs. To further elucidate the physiological significance of the expression of these receptors, we investigated somatostatin-evoked electrophysiological responses and calcium signaling in gonadotrophs, thyrotrophs, somatotrophs, and lactotrophs. Most gonadotrophs and thyrotrophs spontaneously generated action potentials of high-amplitude and short duration. Somatostatin administration silenced both cell types without affecting intracellular calcium concentrations. In contrast, lactotrophs and somatotrophs spontaneously generated low-amplitude, bursting action potentials in conjunction with calcium transients, both of which were silenced by somatostatin. Furthermore, somatostatin inhibited voltage-gated calcium signaling and hormone secretion induced by protein-coupled receptors in all cell types expressing SSTRs, but the inhibition was more pronounced in somatotrophs. The pattern of inhibition of electrical activity and calcium signaling was consistent with direct and indirect inhibition of voltage-gated calcium channels, the latter being triggered by cell type–specific hyperpolarization. These finding suggest that the action of somatostatin in somatotrophs is enhanced by the expression of several types of SSTRs and their slow desensitization. Furthermore, somatostatin may play a role in the electrical resynchronization of gonadotroph, thyrotrophs, and lactotrophs. The lack of SSTR expression in corticotrophs and melanotrophs renders them insensitive to somatostatin, maintaining their excitability and readiness to respond to stress signals [Reference 1].
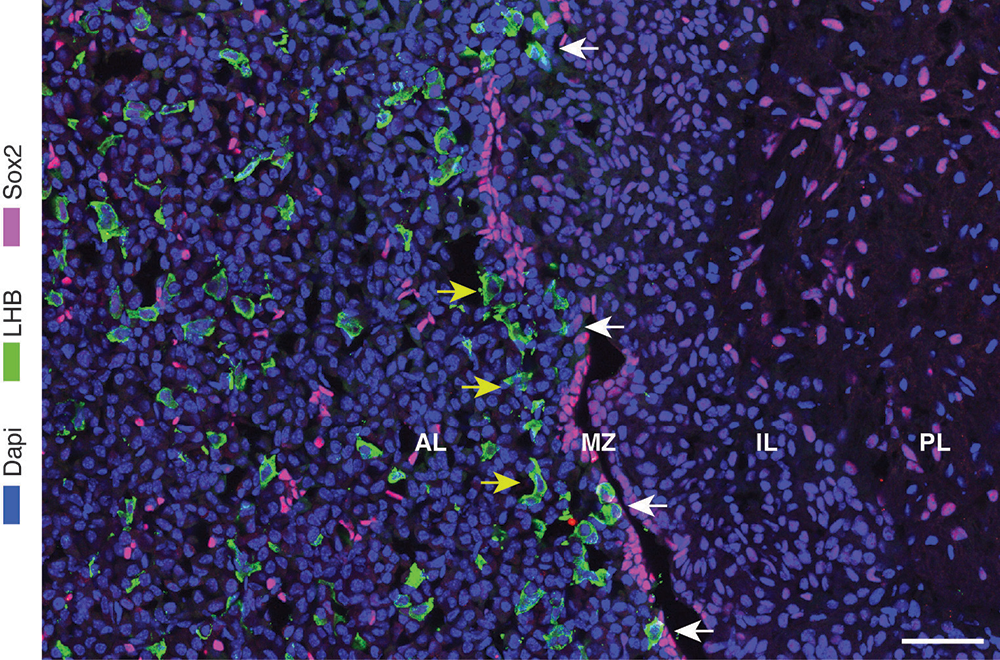
Figure 1. Distribution of Sox2–positive cells and gonadotrophs in the mouse pituitary
Immunostaining of the pituitary sections from 9-week-old female mice: anterior lobe (AL), marginal zone (MZ), intermediate lobe (IL) and posterior lobe (PL). Magenta indicates Sox2–expressing cells, green indicates LHB–positive gonadotrophs, and blue indicates DAPI–labeled cell nuclei.
Figure 1. Distribution of Sox2–positive cells and gonadotrophs in the mouse pituitary
Immunostaining of the pituitary sections from 9-week-old female mice: anterior lobe (AL), marginal zone (MZ), intermediate lobe (IL) and posterior lobe (PL). Magenta indicates Sox2–expressing cells, green indicates LHB–positive gonadotrophs, and blue indicates DAPI–labeled cell nuclei.
Neuroendocrine cell–specific function of Ptprn and Ptprn2 genes
We continue to investigate genes expressed in mammalian hypothalamic and pituitary cells and their roles in cellular signaling and function. Our recent work focused on protein tyrosine phosphatase receptors type N and N2 (PTPRN and PTPRN2) and phosphatidylinositol 4-kinase alpha (PI4KA) genes and their products. PTPRN1 and PTPRN2 are atypical members of the protein tyrosine phosphatase receptors. They are pseudophosphatases resulting from mutation of two residues in the catalytic domain, but they exhibit other cellular functions through a still unknown mechanism. Their genes are expressed in neuroendocrine cells of the hypothalamus and pituitary, but their knockout only affects hypothalamic-pituitary-gonadal and hypothalamic-pituitary-adrenal functions.
Both gonadotropin-releasing hormone (GnRH)–secreting neurons and kisspeptin-secreting neurons also express Ptprn+Ptprn, as do pituitary gonadotrophs, but double knockout (DKO) of these genes did not directly affect the function of GnRH neurons and gonadotrophs. However, kisspeptin staining was significantly reduced in both the arcuate nucleus of female and male DKO mice and in the RP3V region (rostral periventricular area of the third ventricle) of female DKO mice, which controls ovulation. DKO also inhibited the expression of Gnrh1 in GnRH neurons and subsequently Lhb, Fshb (follicle-stimulating hormone subunit beta), and Gnrhr in pituitary gonadotrophs, as well as the accumulation and release of LH (luteinizing hormone). This was accompanied by significant changes in ovarian steroidogenesis and gene expression in DKO females, resulting in the absence of puberty and of the development of female reproductive organs. In contrast, no changes were observed in testicular steroidogenesis and spermatogenesis, nor in seminal vesicles development in DKO males.
In contrast, DKO increased the expression of the corticotroph/melanotroph genes Pomc and Tbx19 and the melanotroph-specific gene Pax7, and the effect was observed in both females and males. We also found in vivo and in vitro increased synthesis and release of beta-endorphin, alpha-MSH (melanocyte stimulating hormone), and ACTH (adrenocorticotropic hormone) in DKO mice, which was associated with increased serum corticosterone levels and adrenal gland mass. DKO also increased the expression of other melanotroph-specific genes, but not of corticotroph-specific genes. Furthermore, hyperplasia of the intermediate lobe was observed in DKO females and males, accompanied by increased POMC immunoreactivity per cell. These results indicate that PTPRNs contribute to hypothalamic-pituitary-adrenal function by being involved in processes that regulate postnatal melanotroph development and Pomc expression.
Thus, although all hypothalamic and pituitary cells express Ptprn and Ptprn2 genes, their DKO only affects the function of specific cells. This is consistent with the hypothesis that PTPRNs act as transcription factors or upstream elements in the control of gene transcription. The cellular specificity of PTPRNs action is consistent with the specificity of promoter activation and repression for different genes, as suggested by the failure of DKO to increase Pomc expression in the hypothalamus but to facilitate its expression in the pituitary [References 2 and 3].
The role of PI4-kinase A in neuroendocrine cell function
PI4-kinase A (PI4KA) is an enzyme that contributes to the production of phosphoinositide PI4P, which serve as a substrate for the production of PI(4,5)P2 and PI(3,4,5)P3. In collaboration with Tamás Balla, we are studying the function of this gene by its specific knockout in hypothalamic and pituitary cells. Knockout of this gene in pituitary gonadotrophs did not alter embryonic establishment, neonatal development, or function of the gonadotroph population. However, during the postnatal period, there was a progressive loss of expression of gonadotroph-specific genes, including Fshb, Lhb, and Gnrhr (gonadotropin-releasing hormone receptor), accompanied by loss of GnRH receptor signaling and reproductive functions in female and male mice. Intracellular administration of inositol-1,4,5-trisphosphate rescued GnRH receptor signaling, indicating that gonadotrophs did not fully dedifferentiate.
We also generated GnRH neuron–specific Pi4ka knockout (KO) to study the role of PI4KA in these cells. KO animals were healthy and indistinguishable from their control littermates in size. However, they were infertile and lacked external features normally associated with sexual maturity (vaginal opening in females and preputial separation in males). Immunohistochemistry against GnRH revealed that the hypothalamus of adult KO mice was devoid of immunoreactive GnRH neurons. In contrast, immunohistochemistry against kisspeptin showed a comparable sex-specific pattern of expression in control littermates and KO mice. At three days of age, Gnrh1 expression levels were similar in KO mice and control littermates, but expression of this gene progressively declined in KO mice and was undetectable by 45 days of age. Immunofluorescence against GnRH at 10 days of age revealed morphological changes in GnRH neurons that preceded their disappearance. In contrast, Kiss1 expression was similar in KO mice and control littermates until the onset of puberty, when the lack of sex-steroid regulation in KO mice affected expression of this gene. The Gnrh1–Cre mouse line used to generate KO mice has strong ectopic expression within the lateral septum, as demonstrated by expression of the tdTomato reporter (red fluorescent protein variant). However, these cells did not display abnormal morphology in KO mice and persisted into adulthood. Therefore, the PI4KA–dependent signaling pathway is not required for the migration of GnRH neurons into the hypothalamus, but is required for their postnatal survival, unlike the ectopic cell type. Current experiments aim to characterize this cell type–specific signaling pathway [Reference 5].
Publications
- Distribution and calcium signaling function of somatostatin receptor subtypes in rat pituitary. Cell Calcium 2024 124:102967
- Protein tyrosine phosphatase receptors N and N2 control pituitary melanotroph development and POMC expression. Endocrinology 2024 165(8):bqae076
- What is known and unknown about the role of neuroendocrine genes Ptprn and Ptprn2. Front Endocrinol (Lausanne) 2025 16:1531723
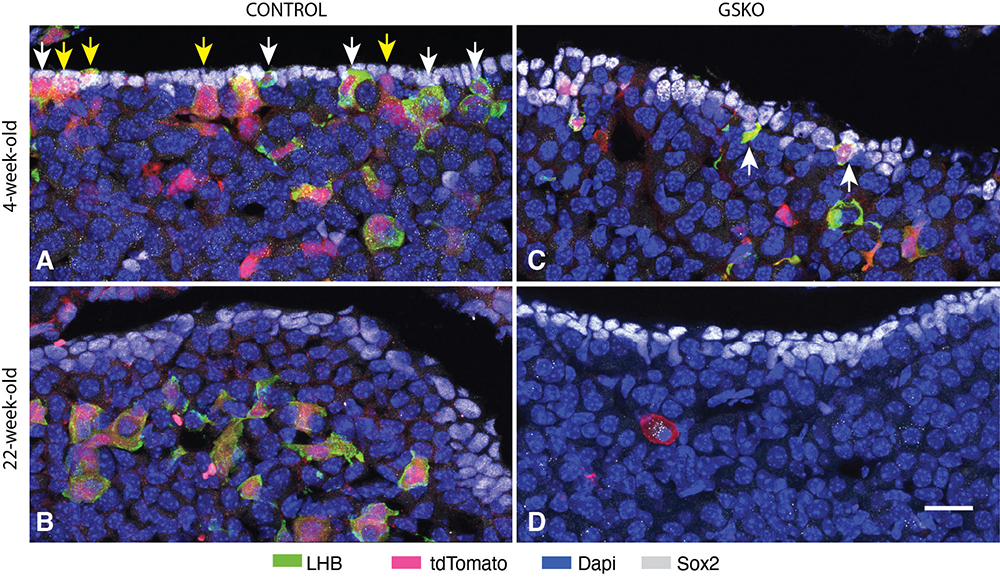
- Heterogeneity of Sox2-expressing cells in mouse pituitary and their roles in postnatal gonadotroph differentiation. Mol Cell Endocrinol 2025 608:112638
- Knockout of PI4-kinase A in GnRH neurons causes their prepubertal death. bioRxiv 2026 submited
Collaborators
- Tamás Balla, MD, PhD, Section on Molecular Signal Transduction, NICHD, Bethesda, MD
- Patrick A. Fletcher, PhD, Laboratory of Biological Modeling, NIDDK, Bethesda, MD
- Arthur Sherman, PhD, Laboratory of Biological Modeling, NIDDK, Bethesda, MD
- Hana Zemkova, PhD, Institute of Physiology of the Czech Academy of Sciences, Prague, Czech Republic
Contact
For more information, email stojilks@mail.nih.gov or visit https://www.nichd.nih.gov/research/atNICHD/Investigators/stojilkovic.